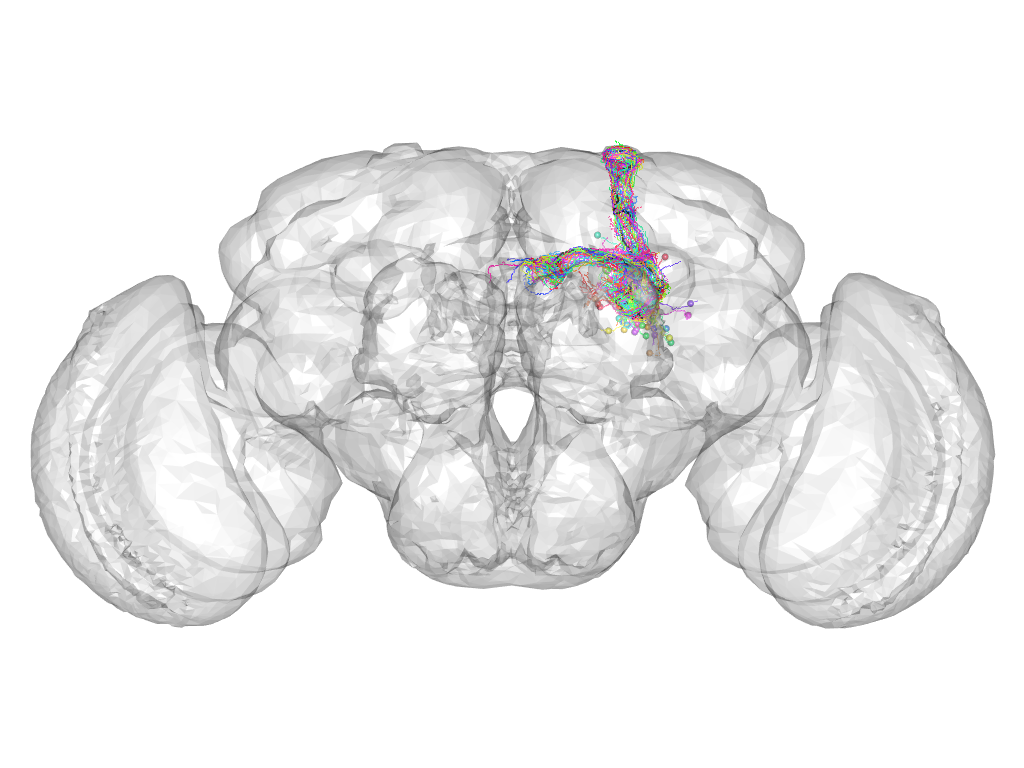
Cluster 101
Part of supercluster XVI
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (fru-M-500143), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 979 | VGlut-F-400141 | XVI | 0.63 |
| 599 | VGlut-F-500640 | XVI | 0.57 |
| 945 | VGlut-F-100136 | XVI | 0.55 |
| 833 | VGlut-F-500352 | XVI | 0.54 |
| 41 | fru-M-700086 | XVI | 0.52 |
| 412 | fru-F-200098 | XVI | 0.51 |
| 96 | fru-M-800049 | XVI | 0.51 |
| 931 | VGlut-F-100079 | XVI | 0.51 |
Cluster composition
This cluster has 38 neurons. The exemplar of this cluster (fru-M-500143) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| fru-M-500143 | 1310 | FruMARCM-M001527_seg003 | fru-Gal4 | M | 7175.21 | 1.00 |
| Gad1-F-700051 | 4084 | GadMARCM-F000673_seg001 | Gad1-Gal4 | F | 4161.04 | 0.53 |
| Cha-F-200353 | 4345 | ChaMARCM-F001402_seg001 | Cha-Gal4 | F | 4144.25 | 0.55 |
| fru-F-500221 | 4663 | FruMARCM-F001826_seg001 | fru-Gal4 | F | 4446.53 | 0.59 |
| fru-F-600112 | 4687 | FruMARCM-F001892_seg001 | fru-Gal4 | F | 4126.42 | 0.56 |
| fru-F-600115 | 4714 | FruMARCM-F002041_seg002 | fru-Gal4 | F | 4604.65 | 0.64 |
| Cha-F-000141 | 5399 | ChaMARCM-F000756_seg002 | Cha-Gal4 | F | 4326.67 | 0.59 |
| Cha-F-600117 | 5438 | ChaMARCM-F000780_seg004 | Cha-Gal4 | F | 4328.41 | 0.59 |
| Cha-F-300210 | 5442 | ChaMARCM-F000782_seg002 | Cha-Gal4 | F | 4414.32 | 0.59 |
| Cha-F-300226 | 5495 | ChaMARCM-F000817_seg001 | Cha-Gal4 | F | 4273.72 | 0.58 |
| Cha-F-000157 | 5585 | ChaMARCM-F000878_seg001 | Cha-Gal4 | F | 4522.36 | 0.63 |
| fru-F-600088 | 6399 | FruMARCM-F001534_seg002 | fru-Gal4 | F | 4430.51 | 0.62 |
| Cha-F-300117 | 7495 | ChaMARCM-F000360_seg001 | Cha-Gal4 | F | 4309.37 | 0.59 |
| Cha-F-800008 | 7504 | ChaMARCM-F000365_seg001 | Cha-Gal4 | F | 3979.76 | 0.60 |
| Cha-F-200083 | 7608 | ChaMARCM-F000453_seg003 | Cha-Gal4 | F | 4311.97 | 0.63 |
| fru-F-500105 | 7759 | FruMARCM-F000661_seg001 | fru-Gal4 | F | 4464.87 | 0.62 |
| fru-F-500124 | 7800 | FruMARCM-F000700_seg001 | fru-Gal4 | F | 4135.47 | 0.56 |
| fru-F-500138 | 7822 | FruMARCM-F000716_seg001 | fru-Gal4 | F | 4385.64 | 0.61 |
| fru-F-000033 | 7894 | FruMARCM-F000825_seg001 | fru-Gal4 | F | 4544.95 | 0.60 |
| VGlut-F-700422 | 8005 | DvGlutMARCM-F003824_seg002 | VGlut-Gal4 | F | 4157.27 | 0.58 |
| VGlut-F-600688 | 8026 | DvGlutMARCM-F003839_seg002 | VGlut-Gal4 | F | 4389.32 | 0.62 |
| Cha-F-600022 | 8339 | ChaMARCM-F000291_seg001 | Cha-Gal4 | F | 4191.22 | 0.63 |
| VGlut-F-600645 | 8922 | DvGlutMARCM-F003498_seg001 | VGlut-Gal4 | F | 4182.77 | 0.57 |
| VGlut-F-300496 | 9173 | DvGlutMARCM-F003247_seg002 | VGlut-Gal4 | F | 4333.75 | 0.57 |
| Gad1-F-600056 | 9477 | GadMARCM-F000373_seg002 | Gad1-Gal4 | F | 4101.88 | 0.56 |
| fru-F-500030 | 9911 | FruMARCM-F000496_seg002 | fru-Gal4 | F | 4476.07 | 0.60 |
| fru-F-600022 | 9941 | FruMARCM-F000526_seg002 | fru-Gal4 | F | 4159.39 | 0.57 |
| Gad1-F-400012 | 10552 | GadMARCM-F000079_seg001 | Gad1-Gal4 | F | 4233.86 | 0.59 |
| Gad1-F-500003 | 10714 | GadMARCM-F000044_seg001 | Gad1-Gal4 | F | 4572.27 | 0.61 |
| VGlut-F-600485 | 11167 | DvGlutMARCM-F003016_seg001 | VGlut-Gal4 | F | 4508.84 | 0.61 |
| fru-F-700018 | 11527 | FruMARCM-F000187_seg001 | fru-Gal4 | F | 4593.15 | 0.61 |
| Trh-F-600076 | 12417 | TPHMARCM-1210F_seg1 | Trh-Gal4 | F | 4573.14 | 0.62 |
| VGlut-F-200350 | 12957 | DvGlutMARCM-F1988_seg1 | VGlut-Gal4 | F | 4597.82 | 0.61 |
| VGlut-F-400168 | 14430 | DvGlutMARCM-F723-X3_seg1 | VGlut-Gal4 | F | 4015.04 | 0.55 |
| VGlut-F-300168 | 14539 | DvGlutMARCM-F808_seg1 | VGlut-Gal4 | F | 4483.28 | 0.58 |
| VGlut-F-500203 | 15251 | DvGlutMARCM-F1571_seg2 | VGlut-Gal4 | F | 2366.40 | 0.46 |
| VGlut-F-100195 | 15419 | DvGlutMARCM-F1776_seg1 | VGlut-Gal4 | F | 4286.02 | 0.57 |
| VGlut-F-400115 | 15501 | DvGlutMARCM-F655-X4_seg4 | VGlut-Gal4 | F | 4631.02 | 0.60 |