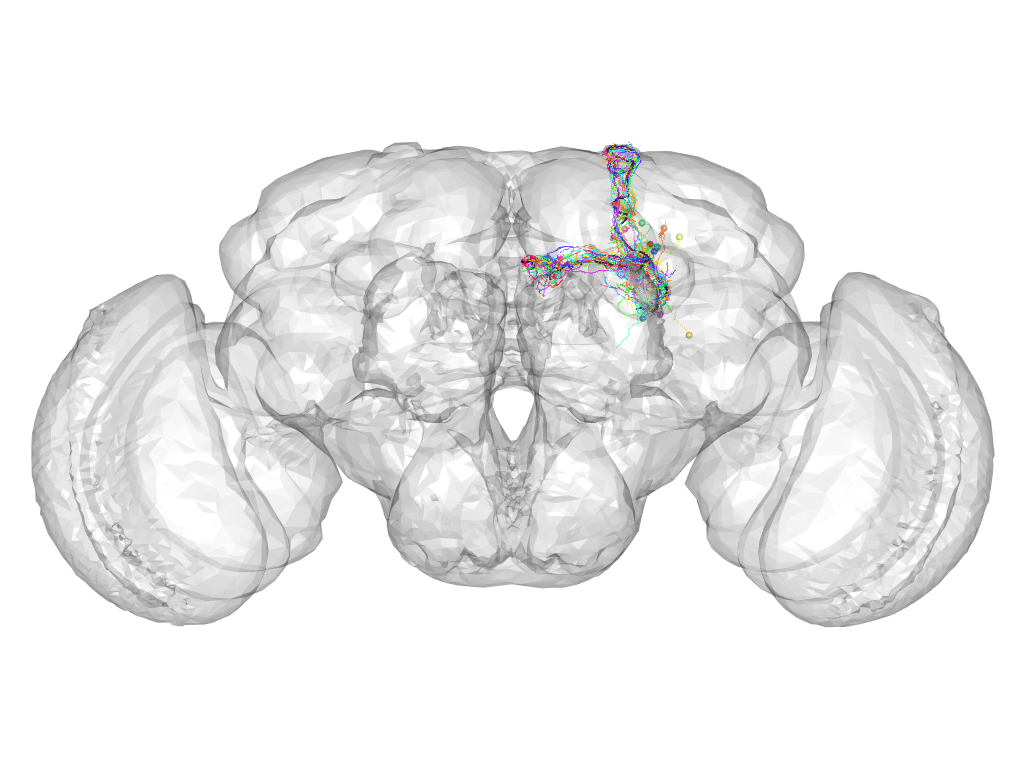
Cluster 599
Part of supercluster XVI
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (VGlut-F-500640), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 979 | VGlut-F-400141 | XVI | 0.58 |
| 816 | VGlut-F-500310 | XVI | 0.56 |
| 101 | fru-M-500143 | XVI | 0.55 |
| 931 | VGlut-F-100079 | XVI | 0.53 |
| 79 | fru-M-300247 | XVI | 0.47 |
Cluster composition
This cluster has 17 neurons. The exemplar of this cluster (VGlut-F-500640) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| VGlut-F-500640 | 9283 | DvGlutMARCM-F003334_seg001 | VGlut-Gal4 | F | 7300.50 | 1.00 |
| fru-M-400108 | 1344 | FruMARCM-M001096_seg001 | fru-Gal4 | M | 4204.15 | 0.57 |
| fru-M-700068 | 1402 | FruMARCM-M000902_seg002 | fru-Gal4 | M | 4350.05 | 0.58 |
| fru-M-400011 | 1795 | FruMARCM-M000323_seg001 | fru-Gal4 | M | 4038.60 | 0.57 |
| fru-M-700034 | 1890 | FruMARCM-M000425_seg001 | fru-Gal4 | M | 4064.77 | 0.57 |
| fru-M-200005 | 2324 | FruMARCM-M000010_seg001 | fru-Gal4 | M | 4499.88 | 0.61 |
| fru-F-600107 | 4650 | FruMARCM-F001814_seg002 | fru-Gal4 | F | 3717.03 | 0.55 |
| VGlut-F-700117 | 6059 | DvGlutMARCM-F002772_seg002 | VGlut-Gal4 | F | 4215.06 | 0.58 |
| Cha-F-600049 | 7537 | ChaMARCM-F000393_seg001 | Cha-Gal4 | F | 4018.15 | 0.54 |
| Cha-F-300070 | 8233 | ChaMARCM-F000230_seg001 | Cha-Gal4 | F | 4135.07 | 0.58 |
| VGlut-F-400772 | 8710 | DvGlutMARCM-F003744_seg001 | VGlut-Gal4 | F | 4294.94 | 0.61 |
| VGlut-F-400717 | 8898 | DvGlutMARCM-F003475_seg001 | VGlut-Gal4 | F | 4306.92 | 0.59 |
| VGlut-F-100284 | 9245 | DvGlutMARCM-F003305_seg001 | VGlut-Gal4 | F | 4197.23 | 0.59 |
| VGlut-F-500644 | 9287 | DvGlutMARCM-F003335_seg003 | VGlut-Gal4 | F | 6433.87 | 0.85 |
| Gad1-F-200111 | 9585 | GadMARCM-F000455_seg001 | Gad1-Gal4 | F | 3988.95 | 0.57 |
| Gad1-F-500005 | 10725 | GadMARCM-F000053_seg001 | Gad1-Gal4 | F | 3604.26 | 0.51 |
| VGlut-F-500063 | 16025 | DvGlutMARCM-F416_seg1 | VGlut-Gal4 | F | 4941.30 | 0.65 |