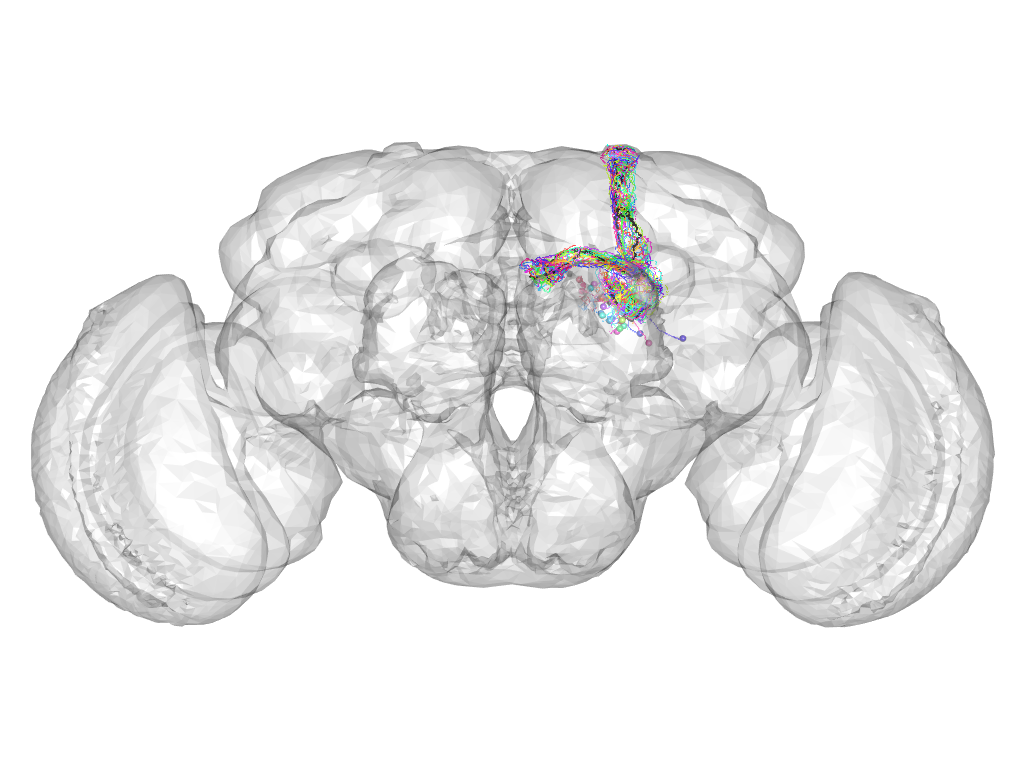
Cluster 945
Part of supercluster XVI
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (VGlut-F-100136), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 96 | fru-M-800049 | XVI | 0.62 |
| 41 | fru-M-700086 | XVI | 0.62 |
| 111 | fru-M-100078 | XVI | 0.59 |
| 412 | fru-F-200098 | XVI | 0.58 |
| 979 | VGlut-F-400141 | XVI | 0.58 |
| 101 | fru-M-500143 | XVI | 0.56 |
| 419 | fru-F-200021 | XVI | 0.51 |
Cluster composition
This cluster has 30 neurons. The exemplar of this cluster (VGlut-F-100136) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| VGlut-F-100136 | 14966 | DvGlutMARCM-F1304_seg1 | VGlut-Gal4 | F | 9589.73 | 1.00 |
| fru-M-400229 | 398 | FruMARCM-M002629_seg002 | fru-Gal4 | M | 5773.90 | 0.65 |
| fru-M-100034 | 1873 | FruMARCM-M000416_seg001 | fru-Gal4 | M | 6222.85 | 0.67 |
| fru-M-500021 | 1913 | FruMARCM-M000468_seg002 | fru-Gal4 | M | 4631.08 | 0.56 |
| Gad1-F-800075 | 4015 | GadMARCM-F000627_seg002 | Gad1-Gal4 | F | 5703.56 | 0.62 |
| fru-F-300098 | 4547 | FruMARCM-F001657_seg001 | fru-Gal4 | F | 6164.94 | 0.67 |
| Cha-F-700171 | 5150 | ChaMARCM-F001198_seg002 | Cha-Gal4 | F | 5491.62 | 0.62 |
| Cha-F-500146 | 5335 | ChaMARCM-F000710_seg001 | Cha-Gal4 | F | 4497.32 | 0.52 |
| Cha-F-300266 | 5693 | ChaMARCM-F000952_seg001 | Cha-Gal4 | F | 5483.85 | 0.61 |
| Cha-F-700004 | 6146 | ChaMARCM-F000028_seg003 | Cha-Gal4 | F | 6112.95 | 0.67 |
| fru-F-200099 | 6342 | FruMARCM-F001494_seg003 | fru-Gal4 | F | 6280.54 | 0.65 |
| fru-F-200102 | 6348 | FruMARCM-F001498_seg002 | fru-Gal4 | F | 6239.45 | 0.65 |
| VGlut-F-600680 | 8006 | DvGlutMARCM-F003825_seg001 | VGlut-Gal4 | F | 6044.79 | 0.66 |
| Cha-F-000024 | 8063 | ChaMARCM-F000134_seg001 | Cha-Gal4 | F | 5672.00 | 0.63 |
| Cha-F-400057 | 8371 | ChaMARCM-F000313_seg001 | Cha-Gal4 | F | 5554.97 | 0.61 |
| Gad1-F-700025 | 8404 | GadMARCM-F000264_seg001 | Gad1-Gal4 | F | 5288.14 | 0.58 |
| VGlut-F-500714 | 8805 | DvGlutMARCM-F003615_seg002 | VGlut-Gal4 | F | 5234.19 | 0.56 |
| VGlut-F-100316 | 8859 | DvGlutMARCM-F003648_seg001 | VGlut-Gal4 | F | 6138.26 | 0.67 |
| Gad1-F-500073 | 9464 | GadMARCM-F000365_seg001 | Gad1-Gal4 | F | 5903.76 | 0.59 |
| Gad1-F-800060 | 9696 | GadMARCM-F000543_seg001 | Gad1-Gal4 | F | 5063.55 | 0.58 |
| fru-F-600010 | 9789 | FruMARCM-F000308_seg001 | fru-Gal4 | F | 5669.02 | 0.61 |
| fru-F-500013 | 9842 | FruMARCM-F000409_seg001 | fru-Gal4 | F | 5398.48 | 0.55 |
| fru-F-600023 | 9942 | FruMARCM-F000527_seg001 | fru-Gal4 | F | 6102.90 | 0.65 |
| Cha-F-500033 | 10180 | ChaMARCM-F000109_seg001 | Cha-Gal4 | F | 5507.29 | 0.60 |
| Gad1-F-200026 | 10732 | GadMARCM-F000057_seg002 | Gad1-Gal4 | F | 5780.51 | 0.62 |
| VGlut-F-900058 | 11063 | DvGlutMARCM-F002940_seg001 | VGlut-Gal4 | F | 6375.49 | 0.67 |
| Gad1-F-300089 | 11387 | GadMARCM-F000340_seg001 | Gad1-Gal4 | F | 5502.57 | 0.59 |
| VGlut-F-500467 | 12235 | DvGlutMARCM-F002698_seg001 | VGlut-Gal4 | F | 5733.87 | 0.59 |
| Gad1-F-300014 | 12629 | GadMARCM-F000033_seg001 | Gad1-Gal4 | F | 5152.10 | 0.54 |
| Gad1-F-200000 | 13164 | GadMARCM-F001_seg1 | Gad1-Gal4 | F | 5616.85 | 0.56 |