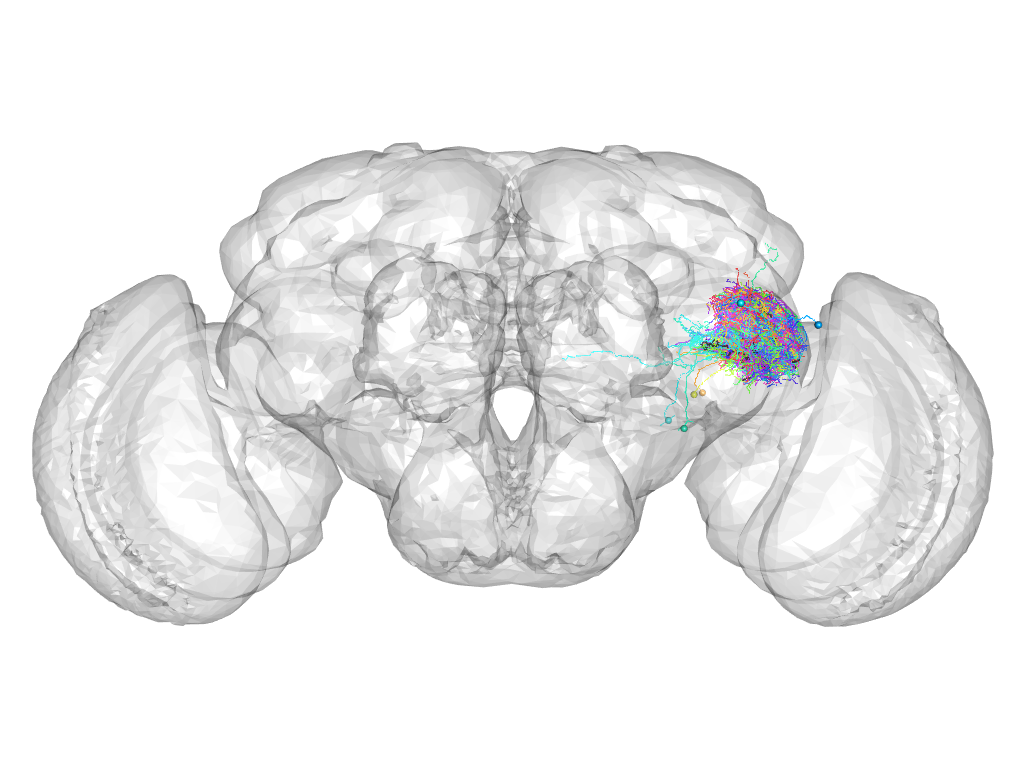
Cluster 895
Part of supercluster X
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (Trh-F-600027), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 939 | VGlut-F-400341 | X | 0.79 |
| 140 | Trh-M-900037 | X | 0.76 |
| 321 | Cha-F-600151 | X | 0.73 |
| 962 | VGlut-F-600088 | X | 0.69 |
| 876 | Trh-F-100023 | X | 0.66 |
| 145 | Trh-M-200079 | VII | 0.58 |
| 171 | Trh-M-000046 | X | 0.55 |
| 200 | TH-M-000060 | X | 0.54 |
| 1038 | VGlut-F-000111 | X | 0.54 |
Cluster composition
This cluster has 73 neurons. The exemplar of this cluster (Trh-F-600027) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| Trh-F-600027 | 14160 | TPHMARCM-587F_seg2 | Trh-Gal4 | F | 28780.58 | 1.00 |
| fru-M-700150 | 3 | FruMARCM-M002278_seg001 | fru-Gal4 | M | 19357.64 | 0.70 |
| fru-M-300256 | 122 | FruMARCM-M002317_seg001 | fru-Gal4 | M | 17902.86 | 0.65 |
| fru-M-700141 | 918 | FruMARCM-M002180_seg001 | fru-Gal4 | M | 18466.34 | 0.68 |
| fru-M-500113 | 1025 | FruMARCM-M001205_seg003 | fru-Gal4 | M | 17961.25 | 0.68 |
| Trh-M-600107 | 2033 | TPHMARCM-M001697_seg002 | Trh-Gal4 | M | 18655.75 | 0.70 |
| Trh-M-600083 | 2096 | TPHMARCM-M001595_seg001 | Trh-Gal4 | M | 18050.59 | 0.69 |
| Trh-M-600084 | 2097 | TPHMARCM-M001596_seg001 | Trh-Gal4 | M | 17964.21 | 0.68 |
| Trh-M-600090 | 2143 | TPHMARCM-M001633_seg001 | Trh-Gal4 | M | 17634.21 | 0.62 |
| Trh-M-700074 | 2596 | TPHMARCM-1186M_seg1 | Trh-Gal4 | M | 15906.57 | 0.63 |
| Trh-M-600066 | 2609 | TPHMARCM-1315M_seg2 | Trh-Gal4 | M | 17199.94 | 0.65 |
| Trh-M-600070 | 2622 | TPHMARCM-M001356_seg001 | Trh-Gal4 | M | 18051.02 | 0.67 |
| Trh-M-500100 | 2679 | TPHMARCM-M001445_seg001 | Trh-Gal4 | M | 19433.49 | 0.70 |
| Trh-M-600077 | 2729 | TPHMARCM-M001494_seg001 | Trh-Gal4 | M | 17084.75 | 0.68 |
| Trh-M-700066 | 2838 | TPHMARCM-1064M_seg1 | Trh-Gal4 | M | 18275.69 | 0.67 |
| Trh-M-700067 | 2839 | TPHMARCM-1065M_seg1 | Trh-Gal4 | M | 18834.55 | 0.71 |
| Trh-M-400071 | 2842 | TPHMARCM-1067M_seg2 | Trh-Gal4 | M | 19622.44 | 0.70 |
| Trh-M-500064 | 2934 | TPHMARCM-869M_seg1 | Trh-Gal4 | M | 18719.67 | 0.67 |
| Trh-M-900011 | 2939 | TPHMARCM-878M_seg2 | Trh-Gal4 | M | 18141.98 | 0.63 |
| Trh-M-700015 | 2981 | TPHMARCM-918M_seg3 | Trh-Gal4 | M | 18196.52 | 0.64 |
| Trh-M-600039 | 2993 | TPHMARCM-936M_seg1 | Trh-Gal4 | M | 19571.81 | 0.71 |
| Trh-M-600041 | 2995 | TPHMARCM-938M_seg1 | Trh-Gal4 | M | 18956.66 | 0.69 |
| Trh-M-600045 | 3002 | TPHMARCM-945M-01_seg1 | Trh-Gal4 | M | 19769.29 | 0.69 |
| Trh-M-700022 | 3025 | TPHMARCM-977M_seg1 | Trh-Gal4 | M | 17185.64 | 0.64 |
| Trh-M-500013 | 3487 | TPHMARCM-360M_seg1 | Trh-Gal4 | M | 17024.97 | 0.66 |
| Trh-M-600003 | 3544 | TPHMARCM-455M_seg1 | Trh-Gal4 | M | 16886.16 | 0.63 |
| Trh-M-600005 | 3546 | TPHMARCM-456M_seg2 | Trh-Gal4 | M | 19196.12 | 0.69 |
| Trh-M-600009 | 3583 | TPHMARCM-520M_seg2 | Trh-Gal4 | M | 19295.51 | 0.70 |
| Trh-M-600011 | 3589 | TPHMARCM-533M_seg2 | Trh-Gal4 | M | 18305.90 | 0.68 |
| Trh-M-500035 | 3623 | TPHMARCM-597M_seg1 | Trh-Gal4 | M | 18808.89 | 0.68 |
| Trh-M-500041 | 3645 | TPHMARCM-660M_seg1 | Trh-Gal4 | M | 19766.33 | 0.69 |
| Trh-M-500045 | 3657 | TPHMARCM-672M_seg1 | Trh-Gal4 | M | 17990.14 | 0.68 |
| Trh-M-500047 | 3663 | TPHMARCM-690M_seg1 | Trh-Gal4 | M | 17867.78 | 0.65 |
| Trh-M-500048 | 3664 | TPHMARCM-704M_seg1 | Trh-Gal4 | M | 18856.93 | 0.68 |
| Gad1-F-200148 | 4100 | GadMARCM-F000689_seg001 | Gad1-Gal4 | F | 17963.18 | 0.60 |
| fru-F-600098 | 4578 | FruMARCM-F001694_seg001 | fru-Gal4 | F | 19422.79 | 0.68 |
| Trh-F-900011 | 5997 | TPHMARCM-711F_seg2 | Trh-Gal4 | F | 18852.27 | 0.70 |
| Cha-F-100044 | 7615 | ChaMARCM-F000458_seg002 | Cha-Gal4 | F | 18856.62 | 0.60 |
| Cha-F-400118 | 7739 | ChaMARCM-F000546_seg001 | Cha-Gal4 | F | 16049.41 | 0.61 |
| VGlut-F-600591 | 9292 | DvGlutMARCM-F003340_seg001 | VGlut-Gal4 | F | 18091.65 | 0.68 |
| Trh-F-700044 | 10452 | TPHMARCM-875F_seg1 | Trh-Gal4 | F | 18163.32 | 0.66 |
| fru-F-800002 | 11443 | FruMARCM-F000063_seg002 | fru-Gal4 | F | 15411.67 | 0.58 |
| Trh-F-600089 | 12441 | TPHMARCM-1235F_seg1 | Trh-Gal4 | F | 17799.67 | 0.66 |
| Trh-F-500131 | 12466 | TPHMARCM-1256F_seg1 | Trh-Gal4 | F | 19110.98 | 0.66 |
| Trh-F-500151 | 12488 | TPHMARCM-1275F_seg2 | Trh-Gal4 | F | 20186.83 | 0.70 |
| Trh-F-500153 | 12497 | TPHMARCM-1282F_seg1 | Trh-Gal4 | F | 18150.92 | 0.66 |
| Trh-F-500172 | 12528 | TPHMARCM-1310F_seg1 | Trh-Gal4 | F | 19012.76 | 0.68 |
| Trh-F-700049 | 12725 | TPHMARCM-905F_seg1 | Trh-Gal4 | F | 17905.73 | 0.65 |
| Trh-F-600067 | 12729 | TPHMARCM-930F_seg2 | Trh-Gal4 | F | 18192.09 | 0.66 |
| Trh-F-500104 | 12731 | TPHMARCM-950F_seg1 | Trh-Gal4 | F | 19613.50 | 0.71 |
| Trh-F-500082 | 13247 | TPHMARCM-734F_seg1 | Trh-Gal4 | F | 18083.12 | 0.69 |
| Trh-F-900015 | 13258 | TPHMARCM-746F_seg2 | Trh-Gal4 | F | 19290.80 | 0.69 |
| Trh-F-400077 | 13269 | TPHMARCM-757F_seg1 | Trh-Gal4 | F | 18000.36 | 0.65 |
| Trh-F-400009 | 13845 | TPHMARCM-114F_seg1 | Trh-Gal4 | F | 19617.92 | 0.72 |
| Trh-F-500003 | 13850 | TPHMARCM-122F_seg1 | Trh-Gal4 | F | 18803.97 | 0.69 |
| Trh-F-500017 | 13864 | TPHMARCM-132F_seg1 | Trh-Gal4 | F | 19357.98 | 0.69 |
| Trh-F-500019 | 13867 | TPHMARCM-137F_seg1 | Trh-Gal4 | F | 17710.82 | 0.58 |
| Trh-F-600001 | 13896 | TPHMARCM-161F_seg1 | Trh-Gal4 | F | 18828.11 | 0.68 |
| Trh-F-600002 | 13897 | TPHMARCM-162F_seg1 | Trh-Gal4 | F | 19660.93 | 0.70 |
| Trh-F-400039 | 13996 | TPHMARCM-286F_seg1 | Trh-Gal4 | F | 17427.78 | 0.65 |
| Trh-F-700001 | 14031 | TPHMARCM-316F_seg1 | Trh-Gal4 | F | 19005.29 | 0.67 |
| Trh-F-500032 | 14051 | TPHMARCM-378F_seg1 | Trh-Gal4 | F | 18751.61 | 0.68 |
| Trh-F-500037 | 14084 | TPHMARCM-433F_seg1 | Trh-Gal4 | F | 18160.10 | 0.68 |
| Trh-F-600012 | 14101 | TPHMARCM-461F_seg1 | Trh-Gal4 | F | 19351.50 | 0.68 |
| Trh-F-600016 | 14107 | TPHMARCM-466F_seg1 | Trh-Gal4 | F | 17297.45 | 0.66 |
| Trh-F-500047 | 14135 | TPHMARCM-525F_seg1 | Trh-Gal4 | F | 18111.65 | 0.69 |
| Trh-F-600025 | 14158 | TPHMARCM-586F_seg1 | Trh-Gal4 | F | 18574.87 | 0.68 |
| Trh-F-300083 | 14169 | TPHMARCM-607F_seg1 | Trh-Gal4 | F | 18434.37 | 0.68 |
| Trh-F-500058 | 14194 | TPHMARCM-636F_seg1 | Trh-Gal4 | F | 19283.88 | 0.67 |
| Trh-F-600031 | 14203 | TPHMARCM-644F_seg1 | Trh-Gal4 | F | 19178.02 | 0.69 |
| Trh-F-600034 | 14208 | TPHMARCM-649F_seg1 | Trh-Gal4 | F | 17338.20 | 0.65 |
| Trh-F-500062 | 14213 | TPHMARCM-653F_seg1 | Trh-Gal4 | F | 19131.65 | 0.69 |
| Trh-F-500063 | 14214 | TPHMARCM-653F_seg2 | Trh-Gal4 | F | 16556.48 | 0.61 |