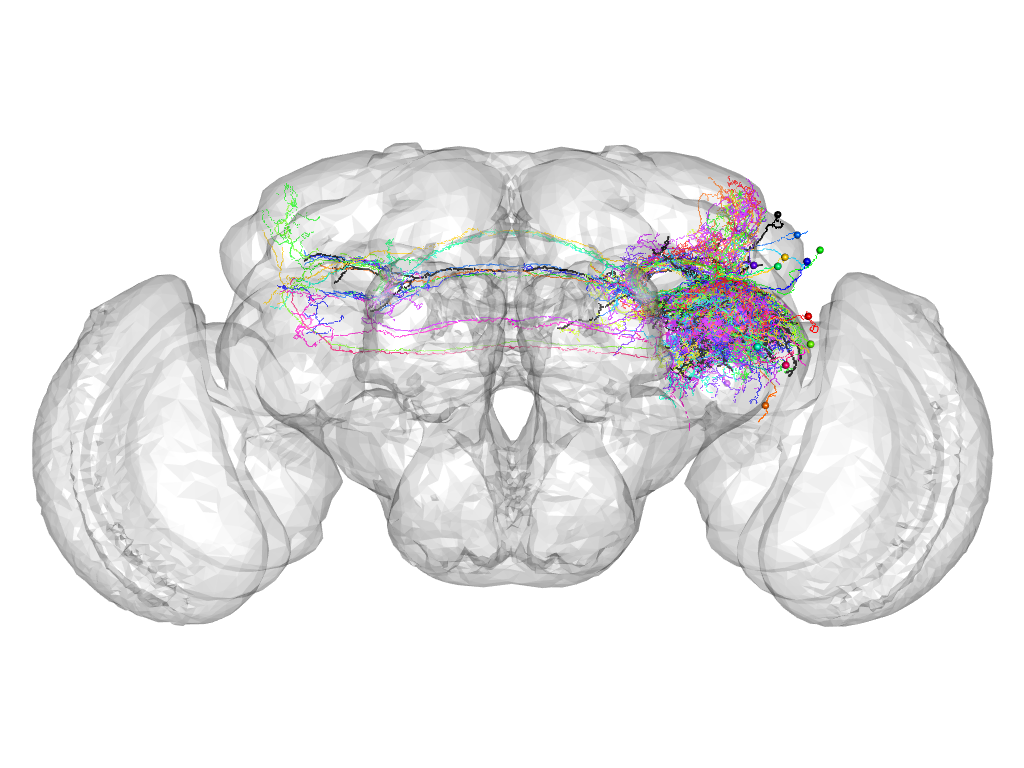
Cluster 1038
Part of supercluster X
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (VGlut-F-000111), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 750 | VGlut-F-400641 | X | 0.60 |
| 939 | VGlut-F-400341 | X | 0.58 |
| 1031 | VGlut-F-400089 | X | 0.56 |
| 865 | TH-F-000025 | X | 0.49 |
| 876 | Trh-F-100023 | X | 0.49 |
Cluster composition
This cluster has 16 neurons. The exemplar of this cluster (VGlut-F-000111) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| VGlut-F-000111 | 16102 | DvGlutMARCM-F485-x2_seg2 | VGlut-Gal4 | F | 77150.64 | 1.00 |
| Trh-M-100027 | 3420 | TPHMARCM-107M_seg1 | Trh-Gal4 | M | 27312.89 | 0.46 |
| Cha-F-000329 | 4131 | ChaMARCM-F001392_seg001 | Cha-Gal4 | F | 31679.55 | 0.48 |
| Cha-F-200137 | 5311 | ChaMARCM-F000692_seg002 | Cha-Gal4 | F | 33713.36 | 0.49 |
| Cha-F-400172 | 5396 | ChaMARCM-F000754_seg002 | Cha-Gal4 | F | 37107.01 | 0.55 |
| Cha-F-300228 | 5497 | ChaMARCM-F000817_seg003 | Cha-Gal4 | F | 38408.23 | 0.51 |
| Cha-F-600041 | 7527 | ChaMARCM-F000376_seg003 | Cha-Gal4 | F | 35584.22 | 0.49 |
| Cha-F-700036 | 8121 | ChaMARCM-F000170_seg001 | Cha-Gal4 | F | 18550.37 | 0.34 |
| Cha-F-500071 | 8291 | ChaMARCM-F000264_seg002 | Cha-Gal4 | F | 43467.47 | 0.57 |
| VGlut-F-000558 | 8857 | DvGlutMARCM-F003646_seg002 | VGlut-Gal4 | F | 36447.62 | 0.58 |
| Gad1-F-300113 | 9580 | GadMARCM-F000450_seg002 | Gad1-Gal4 | F | 51634.19 | 0.69 |
| Cha-F-400011 | 10076 | ChaMARCM-F000036_seg001 | Cha-Gal4 | F | 41616.99 | 0.59 |
| VGlut-F-100199 | 12766 | DvGlutMARCM-F1820_seg1 | VGlut-Gal4 | F | 36569.83 | 0.52 |
| VGlut-F-100125 | 14894 | DvGlutMARCM-F1238_seg1 | VGlut-Gal4 | F | 42005.16 | 0.53 |
| VGlut-F-000328 | 14999 | DvGlutMARCM-F1339_seg1 | VGlut-Gal4 | F | 43036.58 | 0.55 |
| VGlut-F-800030 | 15107 | DvGlutMARCM-F1441_seg1 | VGlut-Gal4 | F | 32235.59 | 0.47 |