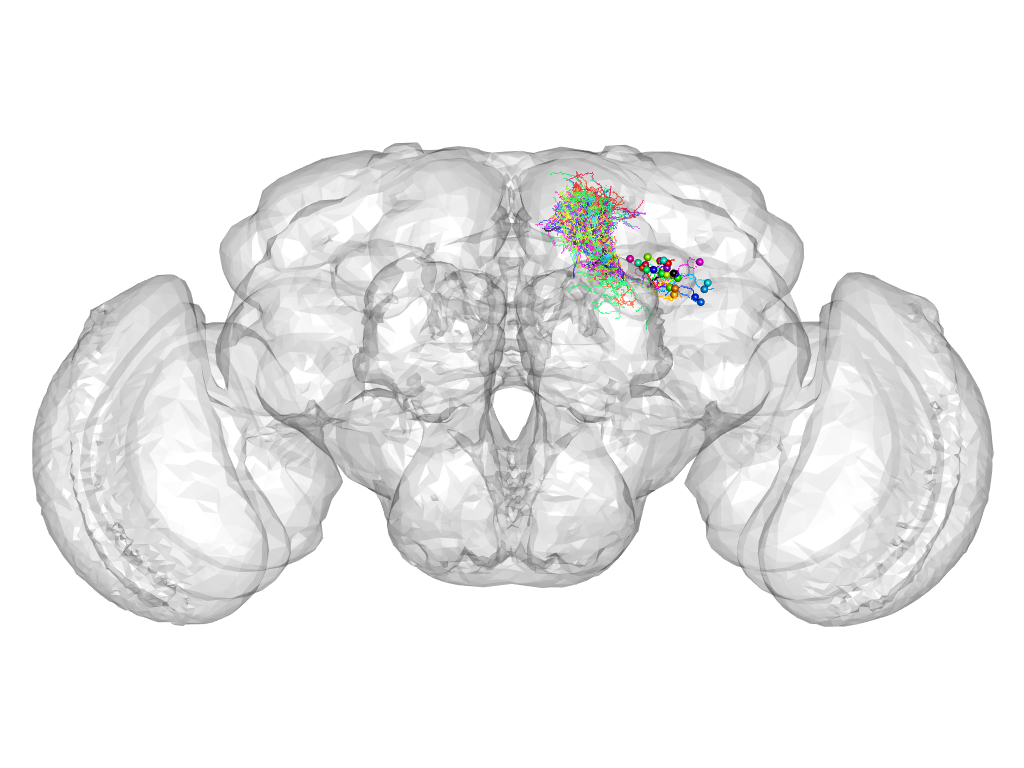
Cluster 182
Part of supercluster XIX
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (Trh-M-600037), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 204 | Trh-M-100025 | XIV | 0.78 |
| 198 | TH-M-000032 | XIX | 0.75 |
| 884 | Trh-F-500028 | XIX | 0.75 |
| 877 | Trh-F-000012 | XIX | 0.72 |
| 190 | TH-M-200014 | XIX | 0.71 |
| 135 | Trh-M-500050 | XIX | 0.71 |
| 994 | VGlut-F-200045 | XIX | 0.68 |
| 42 | fru-M-300180 | XIV | 0.67 |
| 197 | TH-M-200095 | XIX | 0.63 |
| 136 | TH-M-000013 | XVII | 0.62 |
| 1003 | VGlut-F-000024 | XIX | 0.60 |
| 867 | TH-F-100077 | XIX | 0.60 |
| 564 | VGlut-F-000535 | XIX | 0.59 |
| 186 | TH-M-600001 | XIX | 0.59 |
| 1032 | VGlut-F-000077 | XIX | 0.57 |
| 143 | Trh-M-400111 | XIX | 0.55 |
| 941 | VGlut-F-200251 | XIX | 0.53 |
| 871 | TH-F-600003 | XIX | 0.52 |
| 387 | Tdc2-F-000040 | XIV | 0.52 |
Cluster composition
This cluster has 26 neurons. The exemplar of this cluster (Trh-M-600037) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| Trh-M-600037 | 2988 | TPHMARCM-927M_seg1 | Trh-Gal4 | M | 9077.22 | 1.00 |
| Trh-M-800018 | 1322 | TPHMARCM-933M_seg1 | Trh-Gal4 | M | 6923.17 | 0.74 |
| Trh-M-500189 | 2107 | TPHMARCM-M001606_seg001 | Trh-Gal4 | M | 6504.86 | 0.67 |
| Trh-M-500163 | 2544 | TPHMARCM-M001530_seg001 | Trh-Gal4 | M | 6848.83 | 0.75 |
| Trh-M-500096 | 2663 | TPHMARCM-M001427_seg003 | Trh-Gal4 | M | 6921.19 | 0.75 |
| Trh-M-500139 | 2736 | TPHMARCM-M001503_seg001 | Trh-Gal4 | M | 6946.70 | 0.76 |
| Trh-M-800004 | 2779 | TPHMARCM-858M_seg1 | Trh-Gal4 | M | 7089.89 | 0.77 |
| Trh-M-400073 | 2844 | TPHMARCM-1070M_seg1 | Trh-Gal4 | M | 5718.05 | 0.63 |
| Trh-M-400044 | 2997 | TPHMARCM-940M_seg1 | Trh-Gal4 | M | 6623.73 | 0.74 |
| Trh-M-700028 | 3036 | TPHMARCM-989M_seg1 | Trh-Gal4 | M | 6843.00 | 0.74 |
| Cha-F-500184 | 3844 | ChaMARCM-F001518_seg001 | Cha-Gal4 | F | 6913.63 | 0.72 |
| Cha-F-400105 | 7590 | ChaMARCM-F000443_seg001 | Cha-Gal4 | F | 5421.25 | 0.51 |
| VGlut-F-500225 | 8429 | DvGlutMARCM-F1672_seg2 | VGlut-Gal4 | F | 6588.76 | 0.69 |
| Gad1-F-700040 | 9685 | GadMARCM-F000535_seg001 | Gad1-Gal4 | F | 6368.35 | 0.71 |
| Trh-F-500120 | 12439 | TPHMARCM-1233F_seg1 | Trh-Gal4 | F | 6447.53 | 0.72 |
| Trh-F-500132 | 12467 | TPHMARCM-1257F_seg1 | Trh-Gal4 | F | 6263.58 | 0.68 |
| Trh-F-500133 | 12468 | TPHMARCM-1257F_seg2 | Trh-Gal4 | F | 6523.16 | 0.70 |
| Trh-F-500144 | 12479 | TPHMARCM-1263F_seg1 | Trh-Gal4 | F | 6706.21 | 0.73 |
| Trh-F-500169 | 12513 | TPHMARCM-1294F_seg2 | Trh-Gal4 | F | 7094.31 | 0.75 |
| Trh-F-600064 | 12726 | TPHMARCM-928F_seg1 | Trh-Gal4 | F | 7688.87 | 0.84 |
| Trh-F-900033 | 12734 | TPHMARCM-955F_seg1 | Trh-Gal4 | F | 6776.60 | 0.72 |
| Trh-F-500090 | 13279 | TPHMARCM-766F_seg1 | Trh-Gal4 | F | 6622.35 | 0.69 |
| Trh-F-400007 | 13835 | TPHMARCM-056F_seg1 | Trh-Gal4 | F | 6703.52 | 0.68 |
| Trh-F-300076 | 14147 | TPHMARCM-568F_seg1 | Trh-Gal4 | F | 6572.31 | 0.72 |
| Trh-F-300086 | 14174 | TPHMARCM-611F_seg1 | Trh-Gal4 | F | 6067.70 | 0.68 |
| VGlut-F-400118 | 15504 | DvGlutMARCM-F656-X2_seg2 | VGlut-Gal4 | F | 6090.64 | 0.63 |