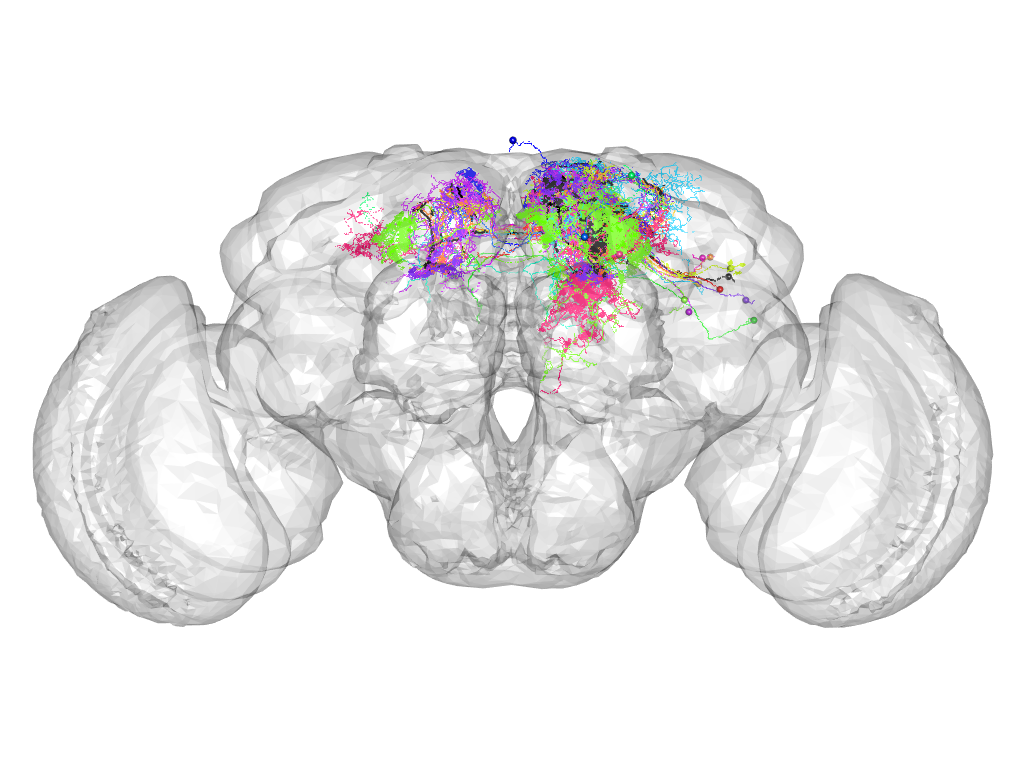
Cluster 197
Part of supercluster XIX
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (TH-M-200095), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 198 | TH-M-000032 | XIX | 0.70 |
| 204 | Trh-M-100025 | XIV | 0.69 |
| 941 | VGlut-F-200251 | XIX | 0.63 |
| 864 | TH-F-100064 | XIX | 0.62 |
| 1003 | VGlut-F-000024 | XIX | 0.61 |
| 859 | TH-F-200089 | XIV | 0.60 |
| 1032 | VGlut-F-000077 | XIX | 0.59 |
| 143 | Trh-M-400111 | XIX | 0.58 |
| 564 | VGlut-F-000535 | XIX | 0.57 |
| 36 | fru-M-200340 | XIX | 0.55 |
| 136 | TH-M-000013 | XVII | 0.52 |
Cluster composition
This cluster has 16 neurons. The exemplar of this cluster (TH-M-200095) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| TH-M-200095 | 3322 | THMARCM-569M_seg | TH-Gal4 | M | 162865.99 | 1.00 |
| TH-M-100001 | 3074 | THMARCM-015M_seg1 | TH-Gal4 | M | 92271.57 | 0.63 |
| TH-M-200010 | 3169 | THMARCM-257M_seg1 | TH-Gal4 | M | 123031.58 | 0.75 |
| TH-M-300034 | 3220 | THMARCM-348M_seg3 | TH-Gal4 | M | 98049.58 | 0.67 |
| TH-M-200087 | 3295 | THMARCM-498M_seg1 | TH-Gal4 | M | 122889.56 | 0.74 |
| TH-M-300055 | 3309 | THMARCM-541M_seg1 | TH-Gal4 | M | 100266.25 | 0.54 |
| Cha-F-000123 | 5344 | ChaMARCM-F000716_seg001 | Cha-Gal4 | F | 87743.11 | 0.50 |
| Cha-F-000139 | 5397 | ChaMARCM-F000755_seg001 | Cha-Gal4 | F | 105758.46 | 0.65 |
| Cha-F-600030 | 7459 | ChaMARCM-F000339_seg001 | Cha-Gal4 | F | 75056.85 | 0.46 |
| fru-F-500102 | 7750 | FruMARCM-F000628_seg001 | fru-Gal4 | F | 105478.36 | 0.59 |
| Gad1-F-900001 | 10612 | GadMARCM-F000120_seg001 | Gad1-Gal4 | F | 61865.51 | 0.50 |
| VGlut-F-600213 | 11779 | DvGlutMARCM-F002347_seg001 | VGlut-Gal4 | F | 80149.62 | 0.55 |
| TH-F-200092 | 13569 | THMARCM-481F_seg1 | TH-Gal4 | F | 127587.64 | 0.75 |
| TH-F-100062 | 13602 | THMARCM-538F_seg2 | TH-Gal4 | F | 90006.55 | 0.58 |
| TH-F-200117 | 13700 | THMARCM-722F_seg1 | TH-Gal4 | F | 120827.34 | 0.76 |
| VGlut-F-000000 | 15601 | DvGlutMARCM-F017_seg1 | VGlut-Gal4 | F | 100476.55 | 0.51 |