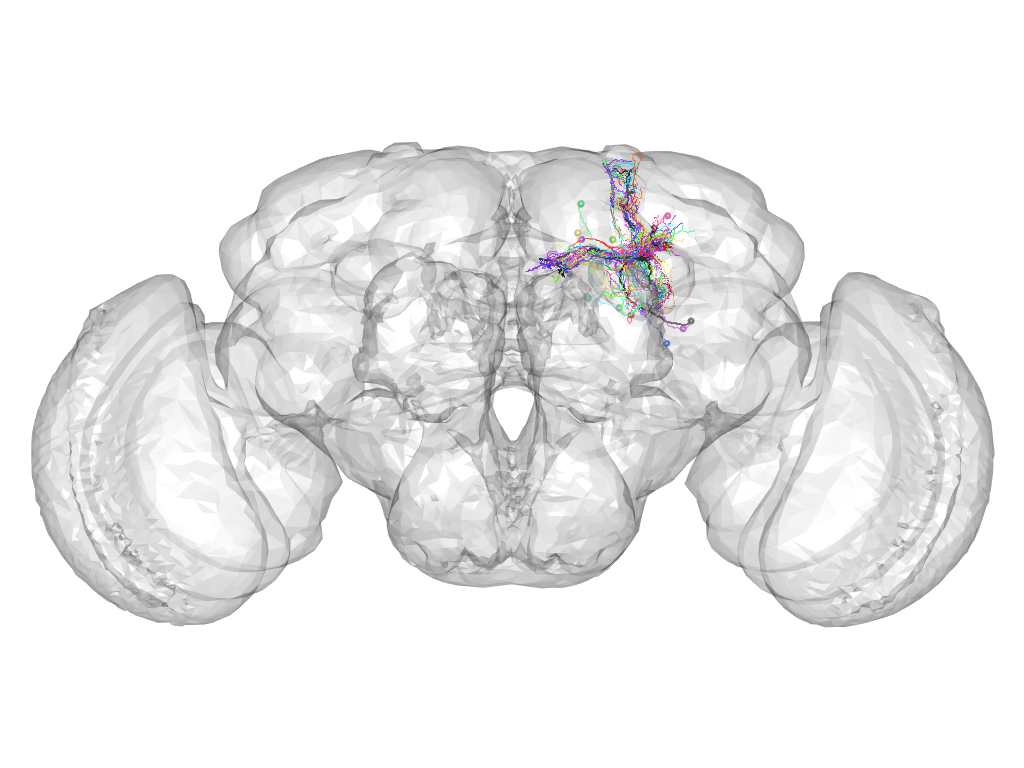
Cluster 833
Part of supercluster XVI
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (VGlut-F-500352), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 816 | VGlut-F-500310 | XVI | 0.54 |
| 979 | VGlut-F-400141 | XVI | 0.48 |
| 169 | Trh-M-200075 | XV | 0.46 |
| 522 | Cha-F-300056 | XVI | 0.46 |
| 599 | VGlut-F-500640 | XVI | 0.46 |
Cluster composition
This cluster has 17 neurons. The exemplar of this cluster (VGlut-F-500352) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| VGlut-F-500352 | 13075 | DvGlutMARCM-F2098_seg1 | VGlut-Gal4 | F | 8109.13 | 1.00 |
| Cha-F-700189 | 3832 | ChaMARCM-F001544_seg002 | Cha-Gal4 | F | 4310.82 | 0.58 |
| Cha-F-300203 | 5374 | ChaMARCM-F000740_seg001 | Cha-Gal4 | F | 3984.36 | 0.51 |
| VGlut-F-700502 | 7043 | DvGlutMARCM-F004175_seg002 | VGlut-Gal4 | F | 3859.32 | 0.46 |
| Cha-F-000038 | 8079 | ChaMARCM-F000144_seg002 | Cha-Gal4 | F | 3733.63 | 0.47 |
| Cha-F-400048 | 8230 | ChaMARCM-F000228_seg001 | Cha-Gal4 | F | 4058.39 | 0.53 |
| Gad1-F-100032 | 9490 | GadMARCM-F000383_seg001 | Gad1-Gal4 | F | 3325.41 | 0.49 |
| Gad1-F-400074 | 9520 | GadMARCM-F000406_seg001 | Gad1-Gal4 | F | 3925.24 | 0.46 |
| Gad1-F-700046 | 9706 | GadMARCM-F000549_seg001 | Gad1-Gal4 | F | 3754.94 | 0.56 |
| Gad1-F-500033 | 10517 | GadMARCM-F000160_seg001 | Gad1-Gal4 | F | 4501.03 | 0.57 |
| Gad1-F-600016 | 10590 | GadMARCM-F000106_seg001 | Gad1-Gal4 | F | 4474.88 | 0.53 |
| Gad1-F-300087 | 11385 | GadMARCM-F000339_seg001 | Gad1-Gal4 | F | 4813.85 | 0.61 |
| Trh-F-500112 | 12407 | TPHMARCM-1201F_seg2 | Trh-Gal4 | F | 4587.05 | 0.56 |
| VGlut-F-400485 | 13004 | DvGlutMARCM-F2032_seg1 | VGlut-Gal4 | F | 4695.19 | 0.56 |
| Gad1-F-000000 | 13180 | GadMARCM-F015_seg1 | Gad1-Gal4 | F | 5394.60 | 0.66 |
| VGlut-F-200231 | 14834 | DvGlutMARCM-F1177_seg1 | VGlut-Gal4 | F | 4609.39 | 0.60 |
| VGlut-F-500240 | 15367 | DvGlutMARCM-F1719_seg1 | VGlut-Gal4 | F | 4046.74 | 0.51 |