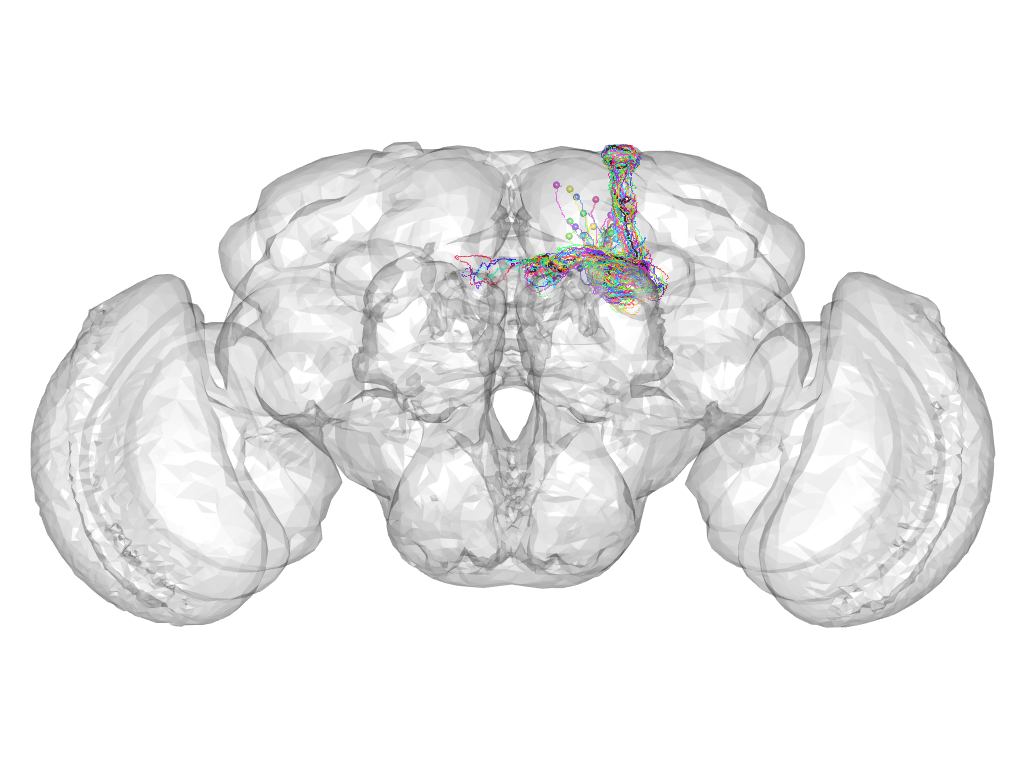
Cluster 522
Part of supercluster XVI
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (Cha-F-300056), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 26 | fru-M-200296 | XVI | 0.60 |
| 533 | Cha-F-900008 | XVI | 0.59 |
| 125 | fru-M-000004 | XVI | 0.58 |
| 407 | fru-F-600073 | XVI | 0.56 |
| 53 | fru-M-000099 | XVI | 0.56 |
| 419 | fru-F-200021 | XVI | 0.55 |
Cluster composition
This cluster has 30 neurons. The exemplar of this cluster (Cha-F-300056) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| Cha-F-300056 | 8187 | ChaMARCM-F000200_seg001 | Cha-Gal4 | F | 7380.22 | 1.00 |
| fru-M-200276 | 99 | FruMARCM-M002297_seg002 | fru-Gal4 | M | 4421.13 | 0.58 |
| fru-M-200207 | 437 | FruMARCM-M001552_seg002 | fru-Gal4 | M | 4887.19 | 0.63 |
| fru-M-500170 | 594 | FruMARCM-M001786_seg003 | fru-Gal4 | M | 5061.73 | 0.67 |
| Cha-F-000111 | 5282 | ChaMARCM-F000674_seg001 | Cha-Gal4 | F | 4250.83 | 0.57 |
| Cha-F-300206 | 5407 | ChaMARCM-F000761_seg001 | Cha-Gal4 | F | 4502.39 | 0.62 |
| Cha-F-000075 | 7239 | ChaMARCM-F000413_seg001 | Cha-Gal4 | F | 4359.24 | 0.57 |
| fru-F-700064 | 7271 | FruMARCM-F000908_seg002 | fru-Gal4 | F | 4204.82 | 0.59 |
| Cha-F-300140 | 7682 | ChaMARCM-F000504_seg003 | Cha-Gal4 | F | 4298.99 | 0.61 |
| Cha-F-500039 | 8034 | ChaMARCM-F000117_seg001 | Cha-Gal4 | F | 4219.23 | 0.58 |
| VGlut-F-500536 | 8454 | DvGlutMARCM-F003052_seg002 | VGlut-Gal4 | F | 3794.37 | 0.50 |
| VGlut-F-600512 | 8544 | DvGlutMARCM-F003117_seg001 | VGlut-Gal4 | F | 4708.59 | 0.60 |
| VGlut-F-500676 | 8939 | DvGlutMARCM-F003507_seg001 | VGlut-Gal4 | F | 4735.23 | 0.66 |
| VGlut-F-500689 | 8952 | DvGlutMARCM-F003515_seg002 | VGlut-Gal4 | F | 4364.64 | 0.59 |
| fru-F-000022 | 9822 | FruMARCM-F000389_seg001 | fru-Gal4 | F | 4493.44 | 0.55 |
| fru-F-500025 | 9906 | FruMARCM-F000493_seg001 | fru-Gal4 | F | 4184.73 | 0.58 |
| fru-F-100028 | 10025 | FruMARCM-F000609_seg001 | fru-Gal4 | F | 4633.33 | 0.61 |
| Cha-F-300021 | 10105 | ChaMARCM-F000059_seg001 | Cha-Gal4 | F | 3675.65 | 0.58 |
| Gad1-F-500010 | 10531 | GadMARCM-F000064_seg001 | Gad1-Gal4 | F | 4776.51 | 0.65 |
| Gad1-F-700011 | 10634 | GadMARCM-F000135_seg001 | Gad1-Gal4 | F | 4247.16 | 0.57 |
| Gad1-F-200017 | 10708 | GadMARCM-F000041_seg001 | Gad1-Gal4 | F | 4828.02 | 0.64 |
| VGlut-F-700217 | 10787 | DvGlutMARCM-F002842_seg001 | VGlut-Gal4 | F | 4049.72 | 0.48 |
| Gad1-F-500044 | 11276 | GadMARCM-F000251_seg001 | Gad1-Gal4 | F | 4596.71 | 0.61 |
| Gad1-F-300075 | 11369 | GadMARCM-F000325_seg001 | Gad1-Gal4 | F | 4292.77 | 0.59 |
| Gad1-F-000029 | 11409 | GadMARCM-F000354_seg002 | Gad1-Gal4 | F | 4309.38 | 0.61 |
| VGlut-F-500383 | 11927 | DvGlutMARCM-F002461_seg001 | VGlut-Gal4 | F | 4476.81 | 0.60 |
| VGlut-F-400479 | 12995 | DvGlutMARCM-F2027_seg1 | VGlut-Gal4 | F | 4734.67 | 0.60 |
| VGlut-F-100233 | 13132 | DvGlutMARCM-F2160_seg1 | VGlut-Gal4 | F | 4920.90 | 0.64 |
| VGlut-F-600143 | 13141 | DvGlutMARCM-F2172_seg1 | VGlut-Gal4 | F | 4592.36 | 0.63 |
| VGlut-F-200268 | 14987 | DvGlutMARCM-F1324_seg1 | VGlut-Gal4 | F | 4553.41 | 0.53 |