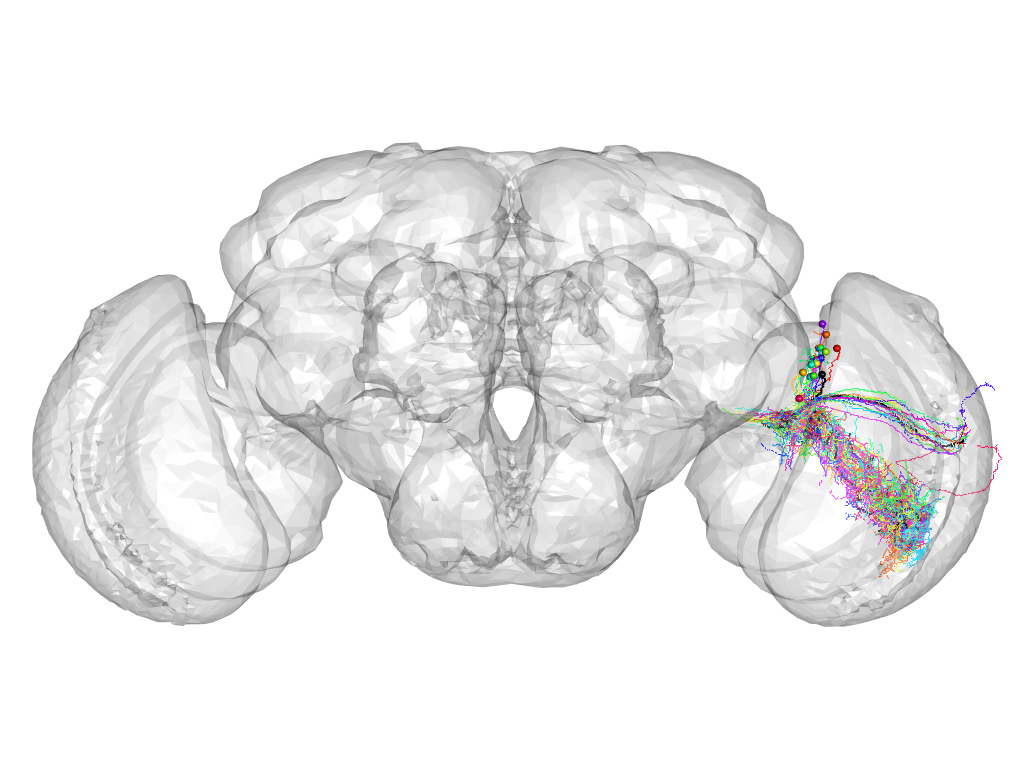
Cluster 150
Part of supercluster III
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (Trh-M-000085), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 208 | Trh-M-500030 | III | 0.63 |
| 661 | Tdc2-F-000027 | VII | 0.62 |
| 4 | fru-M-500226 | III | 0.61 |
| 952 | VGlut-F-600052 | III | 0.58 |
| 76 | fru-M-300240 | III | 0.58 |
| 173 | Trh-M-000051 | III | 0.57 |
| 1006 | VGlut-F-500043 | III | 0.54 |
| 209 | Trh-M-600023 | III | 0.54 |
| 759 | VGlut-F-600257 | III | 0.54 |
| 874 | Trh-F-100011 | III | 0.53 |
Cluster composition
This cluster has 18 neurons. The exemplar of this cluster (Trh-M-000085) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| Trh-M-000085 | 2256 | TPHMARCM-M001743_seg001 | Trh-Gal4 | M | 11981.47 | 1.00 |
| fru-M-600019 | 1762 | FruMARCM-M000250_seg001 | fru-Gal4 | M | 8110.75 | 0.67 |
| Trh-M-000053 | 2059 | TPHMARCM-M001564_seg001 | Trh-Gal4 | M | 8667.04 | 0.67 |
| Trh-M-000059 | 2067 | TPHMARCM-M001570_seg001 | Trh-Gal4 | M | 8278.49 | 0.67 |
| Trh-M-600092 | 2169 | TPHMARCM-M001659_seg001 | Trh-Gal4 | M | 8286.95 | 0.64 |
| Trh-M-000069 | 2216 | TPHMARCM-M001707_seg001 | Trh-Gal4 | M | 6716.05 | 0.57 |
| Trh-M-000086 | 2258 | TPHMARCM-M001745_seg001 | Trh-Gal4 | M | 7595.21 | 0.65 |
| Trh-M-200011 | 2543 | TPHMARCM-193M_seg1 | Trh-Gal4 | M | 8115.58 | 0.63 |
| Trh-M-100016 | 3380 | TPHMARCH-082M_seg1 | Trh-Gal4 | M | 7871.87 | 0.59 |
| Trh-M-000020 | 3493 | TPHMARCM-364M_seg1 | Trh-Gal4 | M | 8054.08 | 0.62 |
| Trh-M-700002 | 3592 | TPHMARCM-536M_seg1 | Trh-Gal4 | M | 8066.02 | 0.60 |
| fru-F-100029 | 10026 | FruMARCM-F000610_seg001 | fru-Gal4 | F | 7892.44 | 0.62 |
| Trh-F-900036 | 11213 | TPHMARCM-F001681_seg001 | Trh-Gal4 | F | 8062.21 | 0.65 |
| fru-F-000003 | 11537 | FruMARCM-F000195_seg001 | fru-Gal4 | F | 8295.55 | 0.64 |
| fru-X-200000 | 11548 | FruMARCM-X000012_seg001 | fru-Gal4 | F | 8219.74 | 0.65 |
| Trh-F-500105 | 12733 | TPHMARCM-954F_seg1 | Trh-Gal4 | F | 8404.52 | 0.67 |
| Trh-F-100014 | 13807 | TPHMARCM-026F_seg1 | Trh-Gal4 | F | 7983.90 | 0.64 |
| Trh-F-300013 | 13893 | TPHMARCM-158F_seg1 | Trh-Gal4 | F | 8677.00 | 0.67 |