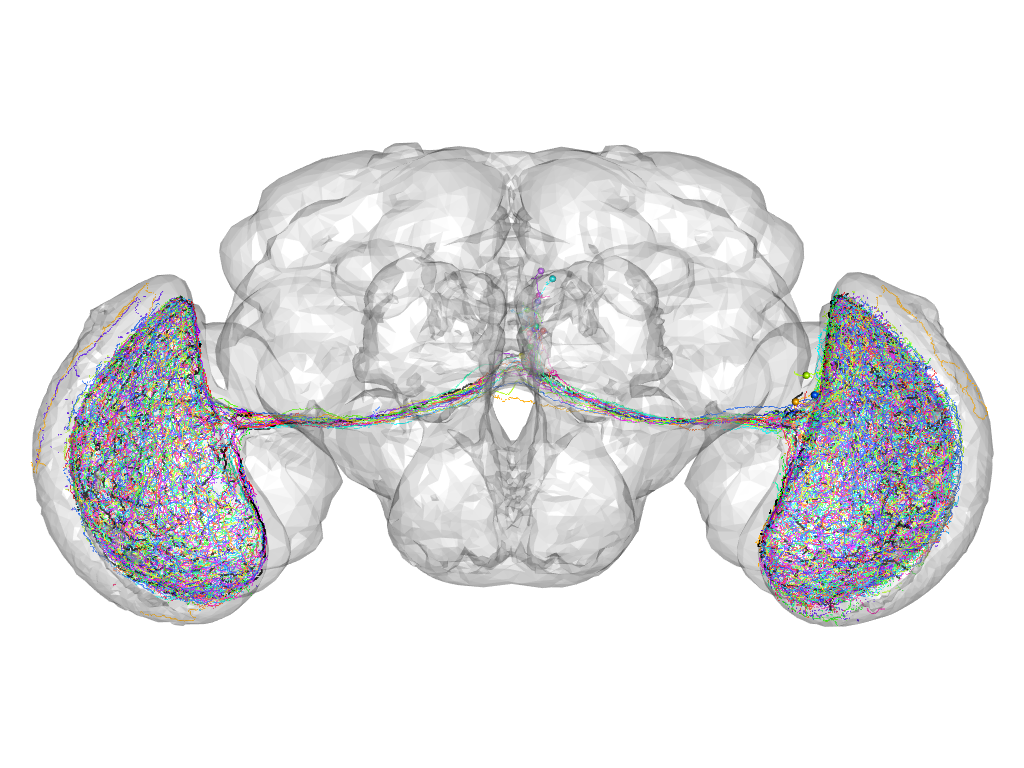
Cluster 173
Part of supercluster III
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (Trh-M-000051), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 468 | VGlut-F-600670 | III | 0.33 |
| 759 | VGlut-F-600257 | III | 0.02 |
| 661 | Tdc2-F-000027 | VII | -0.02 |
| 1006 | VGlut-F-500043 | III | -0.11 |
| 874 | Trh-F-100011 | III | -0.15 |
Cluster composition
This cluster has 19 neurons. The exemplar of this cluster (Trh-M-000051) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| Trh-M-000051 | 2719 | TPHMARCM-M001483_seg001 | Trh-Gal4 | M | 92548.88 | 1.00 |
| Trh-M-000138 | 1730 | TPHMARCM-M001807_seg001 | Trh-Gal4 | M | 69095.92 | 0.75 |
| Trh-M-000154 | 1745 | TPHMARCM-M001822_seg001 | Trh-Gal4 | M | 69110.50 | 0.75 |
| npf-M-300000 | 2029 | npfMARCM-M000006_seg001 | npf-GAL4 | M | -19141.04 | 0.05 |
| Trh-M-000050 | 2718 | TPHMARCM-M001482_seg001 | Trh-Gal4 | M | 69320.27 | 0.75 |
| Cha-F-000019 | 5966 | ChaMARCM-F000097_seg001 | Cha-Gal4 | F | 45506.70 | 0.48 |
| Trh-F-000079 | 10059 | TPHMARCM-F001837_seg001 | Trh-Gal4 | F | 66294.53 | 0.72 |
| Trh-F-000091 | 10069 | TPHMARCM-F001850_seg001 | Trh-Gal4 | F | 66304.56 | 0.72 |
| Trh-F-000042 | 10203 | TPHMARCM-1251_seg001 | Trh-Gal4 | F | 67625.15 | 0.74 |
| Trh-F-000045 | 10205 | TPHMARCM-1322F_seg1 | Trh-Gal4 | F | 69788.85 | 0.75 |
| Trh-F-000015 | 10444 | TPHMARCM-059F_seg1 | Trh-Gal4 | F | 68930.70 | 0.72 |
| VGlut-F-000192 | 10762 | DVGlutMARCMFemale-0521-01-F773_seg1 | VGlut-Gal4 | F | 71002.59 | 0.72 |
| VGlut-F-300437 | 11154 | DvGlutMARCM-F003007_seg001 | VGlut-Gal4 | F | 42084.57 | 0.52 |
| VGlut-F-000429 | 12964 | DvGlutMARCM-F1995-montage_seg1 | VGlut-Gal4 | F | 69466.17 | 0.75 |
| Trh-F-000016 | 13840 | TPHMARCM-104F-montage_seg1 | Trh-Gal4 | F | 69813.63 | 0.71 |
| Trh-F-400043 | 14033 | TPHMARCM-318F-montage_seg1 | Trh-Gal4 | F | 67640.55 | 0.74 |
| Trh-F-000026 | 14034 | TPHMARCM-323F-montage_seg1 | Trh-Gal4 | F | 65189.77 | 0.72 |
| Trh-F-300093 | 14190 | TPHMARCM-632F-montage_seg1 | Trh-Gal4 | F | 67206.97 | 0.73 |
| VGlut-F-200287 | 15198 | DvGlutMARCM-F1526-montage_seg1 | VGlut-Gal4 | F | 67142.63 | 0.73 |