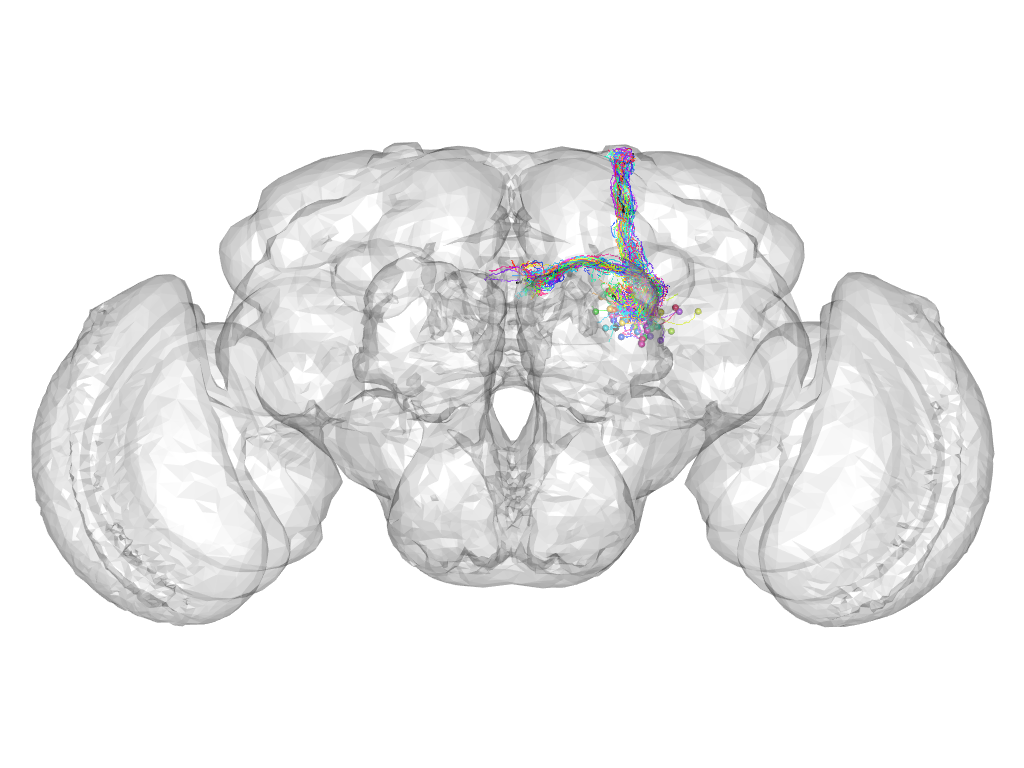
Cluster 111
Part of supercluster XVI
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (fru-M-100078), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 96 | fru-M-800049 | XVI | 0.69 |
| 41 | fru-M-700086 | XVI | 0.68 |
| 412 | fru-F-200098 | XVI | 0.67 |
| 945 | VGlut-F-100136 | XVI | 0.66 |
| 979 | VGlut-F-400141 | XVI | 0.61 |
| 101 | fru-M-500143 | XVI | 0.53 |
Cluster composition
This cluster has 44 neurons. The exemplar of this cluster (fru-M-100078) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| fru-M-100078 | 1500 | FruMARCM-M001051_seg002 | fru-Gal4 | M | 4703.75 | 1.00 |
| fru-M-300202 | 526 | FruMARCM-M001647_seg003 | fru-Gal4 | M | 3225.16 | 0.67 |
| fru-M-300215 | 662 | FruMARCM-M001871_seg003 | fru-Gal4 | M | 3328.72 | 0.69 |
| fru-M-300220 | 681 | FruMARCM-M001885_seg002 | fru-Gal4 | M | 3285.49 | 0.69 |
| fru-M-700109 | 690 | FruMARCM-M001894_seg001 | fru-Gal4 | M | 3370.69 | 0.72 |
| fru-M-500205 | 806 | FruMARCM-M002016_seg001 | fru-Gal4 | M | 3342.93 | 0.68 |
| fru-M-100183 | 830 | FruMARCM-M002033_seg002 | fru-Gal4 | M | 3245.84 | 0.67 |
| fru-M-100186 | 833 | FruMARCM-M002035_seg002 | fru-Gal4 | M | 3483.73 | 0.72 |
| fru-M-500063 | 1369 | FruMARCM-M000878_seg002 | fru-Gal4 | M | 3250.48 | 0.69 |
| fru-M-500090 | 1443 | FruMARCM-M001013_seg001 | fru-Gal4 | M | 3204.78 | 0.66 |
| fru-M-100083 | 1521 | FruMARCM-M001065_seg001 | fru-Gal4 | M | 3328.74 | 0.69 |
| fru-M-100053 | 1584 | FruMARCM-M000652_seg003 | fru-Gal4 | M | 744.23 | 0.42 |
| fru-M-000028 | 1648 | FruMARCM-M000788_seg001 | fru-Gal4 | M | 3217.64 | 0.69 |
| fru-M-000029 | 1653 | FruMARCM-M000792_seg001 | fru-Gal4 | M | 3163.07 | 0.68 |
| fru-M-400010 | 1794 | FruMARCM-M000322_seg001 | fru-Gal4 | M | 3407.40 | 0.73 |
| fru-M-100045 | 1915 | FruMARCM-M000473_seg001 | fru-Gal4 | M | 3216.62 | 0.68 |
| fru-M-200043 | 2455 | FruMARCM-M000214_seg001 | fru-Gal4 | M | 3163.55 | 0.67 |
| fru-M-800008 | 2478 | FruMARCM-M000233_seg002 | fru-Gal4 | M | 3169.97 | 0.66 |
| fru-M-800009 | 2479 | FruMARCM-M000234_seg001 | fru-Gal4 | M | 2363.79 | 0.60 |
| Gad1-F-900051 | 4029 | GadMARCM-F000636_seg002 | Gad1-Gal4 | F | 3311.98 | 0.67 |
| Cha-F-100080 | 5258 | ChaMARCM-F000657_seg001 | Cha-Gal4 | F | 2864.31 | 0.61 |
| Cha-F-000151 | 5544 | ChaMARCM-F000846_seg001 | Cha-Gal4 | F | 3055.91 | 0.63 |
| fru-F-300066 | 6203 | FruMARCM-F001130_seg002 | fru-Gal4 | F | 3367.66 | 0.72 |
| VGlut-F-700461 | 6764 | DvGlutMARCM-F003967_seg002 | VGlut-Gal4 | F | 3338.14 | 0.68 |
| Cha-F-100076 | 7163 | ChaMARCM-F000619_seg002 | Cha-Gal4 | F | 2858.27 | 0.61 |
| fru-F-700079 | 7291 | FruMARCM-F000921_seg001 | fru-Gal4 | F | 3163.58 | 0.67 |
| fru-F-700105 | 7351 | FruMARCM-F000984_seg002 | fru-Gal4 | F | 3251.72 | 0.73 |
| Cha-F-200085 | 7637 | ChaMARCM-F000473_seg001 | Cha-Gal4 | F | 2885.63 | 0.66 |
| Cha-F-300144 | 7729 | ChaMARCM-F000540_seg001 | Cha-Gal4 | F | 2863.33 | 0.58 |
| Cha-F-700024 | 8042 | ChaMARCM-F000121_seg002 | Cha-Gal4 | F | 2755.85 | 0.57 |
| Cha-F-000022 | 8053 | ChaMARCM-F000126_seg001 | Cha-Gal4 | F | 3157.19 | 0.67 |
| Cha-F-900005 | 8092 | ChaMARCM-F000153_seg001 | Cha-Gal4 | F | 3208.01 | 0.66 |
| Cha-F-300077 | 8302 | ChaMARCM-F000271_seg001 | Cha-Gal4 | F | 3055.42 | 0.63 |
| VGlut-F-500589 | 9354 | DvGlutMARCM-F003187_seg001 | VGlut-Gal4 | F | 3202.44 | 0.64 |
| fru-F-300026 | 9787 | FruMARCM-F000306_seg002 | fru-Gal4 | F | 2812.67 | 0.59 |
| fru-F-500020 | 9891 | FruMARCM-F000485_seg001 | fru-Gal4 | F | 3021.61 | 0.63 |
| Cha-F-500018 | 10135 | ChaMARCM-F000080_seg001 | Cha-Gal4 | F | 2840.06 | 0.62 |
| Gad1-F-500015 | 10499 | GadMARCM-F000152_seg002 | Gad1-Gal4 | F | 3086.97 | 0.67 |
| Gad1-F-500037 | 10525 | GadMARCM-F000165_seg002 | Gad1-Gal4 | F | 3094.71 | 0.66 |
| Gad1-F-900003 | 10614 | GadMARCM-F000121_seg001 | Gad1-Gal4 | F | 2889.31 | 0.64 |
| VGlut-F-500485 | 11012 | DvGlutMARCM-F002898_seg001 | VGlut-Gal4 | F | 2898.33 | 0.61 |
| VGlut-F-200443 | 11083 | DvGlutMARCM-F002957_seg002 | VGlut-Gal4 | F | 3305.78 | 0.66 |
| fru-F-400026 | 11492 | FruMARCM-F000157_seg001 | fru-Gal4 | F | 2857.44 | 0.61 |
| VGlut-F-000137 | 16154 | DvGlutMARCM-F536_seg2 | VGlut-Gal4 | F | 3047.45 | 0.63 |