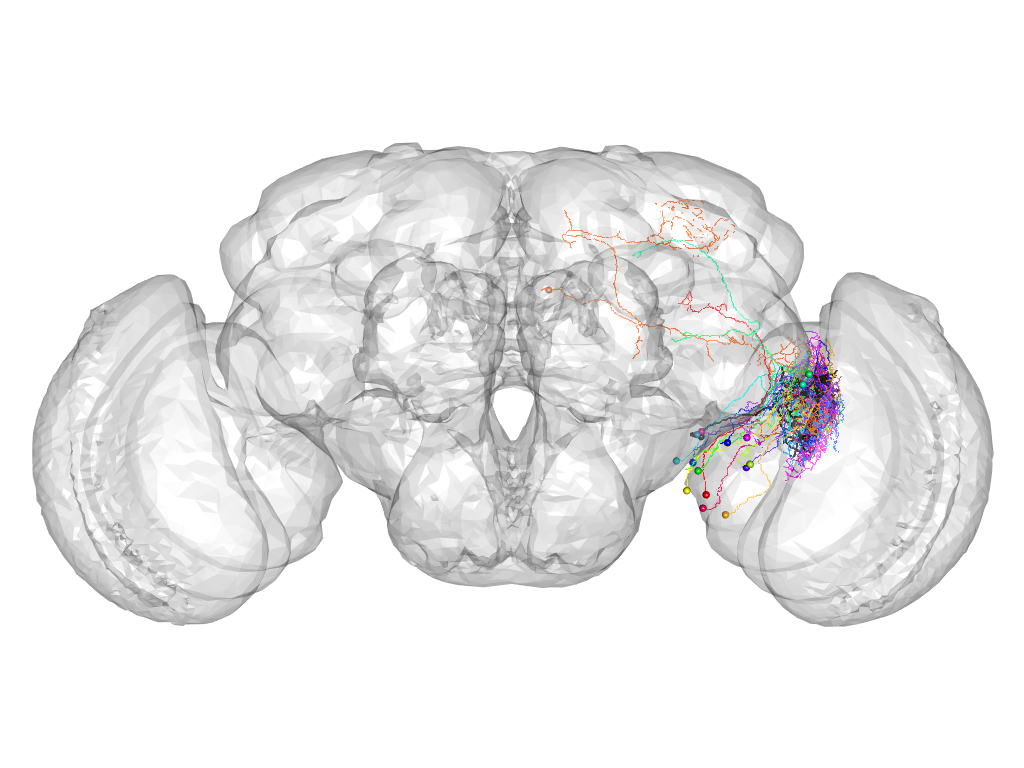
Cluster 224
Part of supercluster I
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (fru-F-800066), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 775 | VGlut-F-500422 | II | 0.77 |
| 977 | VGlut-F-400133 | II | 0.63 |
| 293 | fru-F-000074 | II | 0.62 |
| 13 | fru-M-500232 | I | 0.59 |
| 256 | Cha-F-000296 | I | 0.58 |
| 509 | VGlut-F-200536 | VII | 0.57 |
| 502 | fru-F-000032 | I | 0.50 |
Cluster composition
This cluster has 19 neurons. The exemplar of this cluster (fru-F-800066) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| fru-F-800066 | 3799 | FruMARCM-F002512_seg001 | fru-Gal4 | F | 44326.88 | 1.00 |
| fru-M-500073 | 1387 | FruMARCM-M000892_seg001 | fru-Gal4 | M | 27356.48 | 0.53 |
| fru-M-100008 | 2361 | FruMARCM-M000054_seg001 | fru-Gal4 | M | 31966.32 | 0.43 |
| Cha-F-200336 | 4303 | ChaMARCM-F001369_seg001 | Cha-Gal4 | F | 19233.84 | 0.48 |
| Cha-F-400164 | 5359 | ChaMARCM-F000728_seg001 | Cha-Gal4 | F | 18429.43 | 0.48 |
| VGlut-F-600699 | 7419 | DvGlutMARCM-F003875_seg002 | VGlut-Gal4 | F | 13151.89 | 0.38 |
| VGlut-F-800139 | 9391 | DvGlutMARCM-F003219_seg001 | VGlut-Gal4 | F | 23777.98 | 0.57 |
| VGlut-F-400679 | 9414 | DvGlutMARCM-F003230_seg001 | VGlut-Gal4 | F | 20087.65 | 0.52 |
| Gad1-F-200101 | 9486 | GadMARCM-F000380_seg001 | Gad1-Gal4 | F | 23489.75 | 0.53 |
| Gad1-F-000040 | 9513 | GadMARCM-F000402_seg001 | Gad1-Gal4 | F | 22551.73 | 0.40 |
| Gad1-F-800044 | 9550 | GadMARCM-F000429_seg002 | Gad1-Gal4 | F | 20439.85 | 0.54 |
| Gad1-F-200115 | 9591 | GadMARCM-F000460_seg001 | Gad1-Gal4 | F | 30730.97 | 0.70 |
| VGlut-F-700143 | 10868 | DvGlutMARCM-F002782_seg001 | VGlut-Gal4 | F | 21921.78 | 0.55 |
| Trh-F-300131 | 12680 | TPHMARCM-1016F_seg1 | Trh-Gal4 | F | 28070.79 | 0.67 |
| VGlut-F-500274 | 12792 | DvGlutMARCM-F1840_seg1 | VGlut-Gal4 | F | 19583.48 | 0.57 |
| VGlut-F-000309 | 14784 | DvGlutMARCM-F1133_seg1 | VGlut-Gal4 | F | 26205.77 | 0.63 |
| VGlut-F-800013 | 15803 | DvGlutMARCM-F213_seg1 | VGlut-Gal4 | F | 26928.88 | 0.60 |
| VGlut-F-400039 | 15835 | DvGlutMARCM-F242-X4_seg3 | VGlut-Gal4 | F | 23384.93 | 0.63 |
| VGlut-F-000096 | 16077 | DvGlutMARCM-F462-X2_seg1 | VGlut-Gal4 | F | 22709.52 | 0.55 |