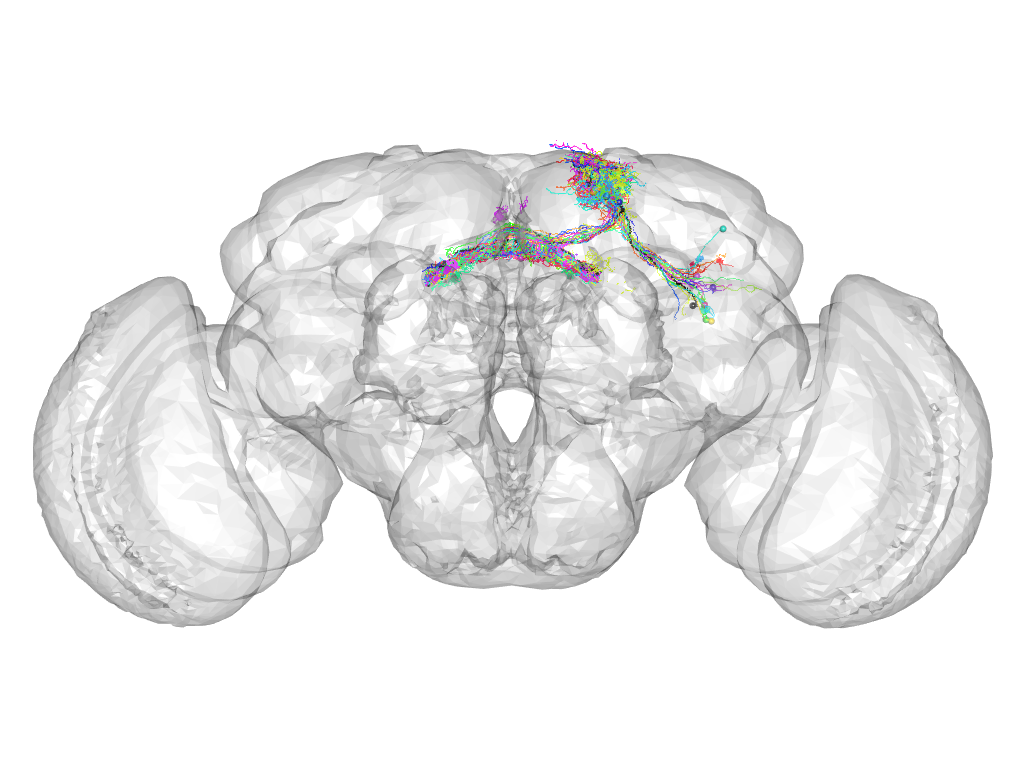
Cluster 887
Part of supercluster XIX
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (Trh-F-100046), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 83 | TH-M-300024 | XIX | 0.73 |
| 141 | Trh-M-400106 | XIX | 0.68 |
| 189 | TH-M-200006 | XIX | 0.50 |
| 753 | VGlut-F-600178 | XIV | 0.45 |
| 193 | TH-M-100039 | XIX | 0.32 |
Cluster composition
This cluster has 22 neurons. The exemplar of this cluster (Trh-F-100046) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| Trh-F-100046 | 14053 | TPHMARCM-379F_seg1 | Trh-Gal4 | F | 16844.67 | 1.00 |
| fru-M-900070 | 5 | FruMARCM-M002507_seg001 | fru-Gal4 | M | 12872.64 | 0.72 |
| fru-M-200205 | 1297 | FruMARCM-M001488_seg001 | fru-Gal4 | M | 12461.68 | 0.71 |
| Trh-M-000093 | 2267 | TPHMARCM-M001754_seg001 | Trh-Gal4 | M | 12536.66 | 0.74 |
| Trh-M-000024 | 2650 | TPHMARCM-M001419_seg001 | Trh-Gal4 | M | 13191.09 | 0.77 |
| Trh-M-000028 | 2654 | TPHMARCM-M001422_seg002 | Trh-Gal4 | M | 12933.42 | 0.74 |
| Trh-M-100080 | 2665 | TPHMARCM-M001429_seg001 | Trh-Gal4 | M | 12915.02 | 0.75 |
| Trh-M-100093 | 2716 | TPHMARCM-M001480_seg001 | Trh-Gal4 | M | 12139.63 | 0.71 |
| Trh-M-300078 | 2944 | TPHMARCM-888M_seg1 | Trh-Gal4 | M | 11903.39 | 0.69 |
| Trh-M-100024 | 3414 | TPHMARCM-099M_seg1 | Trh-Gal4 | M | 12798.06 | 0.74 |
| Trh-M-000014 | 3477 | TPHMARCM-348M_seg1 | Trh-Gal4 | M | 12420.32 | 0.70 |
| Cha-F-100271 | 4230 | ChaMARCM-F001318_seg001 | Cha-Gal4 | F | 11826.92 | 0.68 |
| fru-F-200148 | 4767 | FruMARCM-F002086_seg001 | fru-Gal4 | F | 13203.34 | 0.74 |
| Trh-F-100070 | 12490 | TPHMARCM-1276F_seg1 | Trh-Gal4 | F | 12384.32 | 0.70 |
| Trh-F-100083 | 12589 | TPHMARCM-F001382_seg001 | Trh-Gal4 | F | 12870.72 | 0.75 |
| Trh-F-100015 | 13808 | TPHMARCM-027F_seg1 | Trh-Gal4 | F | 12302.70 | 0.74 |
| Trh-F-100017 | 13810 | TPHMARCM-029F_seg1 | Trh-Gal4 | F | 12673.40 | 0.75 |
| Trh-F-000011 | 13823 | TPHMARCM-044F_seg1 | Trh-Gal4 | F | 12897.51 | 0.75 |
| Trh-F-100027 | 13834 | TPHMARCM-055F_seg1 | Trh-Gal4 | F | 12315.59 | 0.71 |
| Trh-F-900001 | 13910 | TPHMARCM-178F_seg1 | Trh-Gal4 | F | 12606.12 | 0.72 |
| Trh-F-200018 | 13939 | TPHMARCM-231F_seg1 | Trh-Gal4 | F | 12331.11 | 0.74 |
| Trh-F-200053 | 14009 | TPHMARCM-297F_seg1 | Trh-Gal4 | F | 12538.55 | 0.71 |