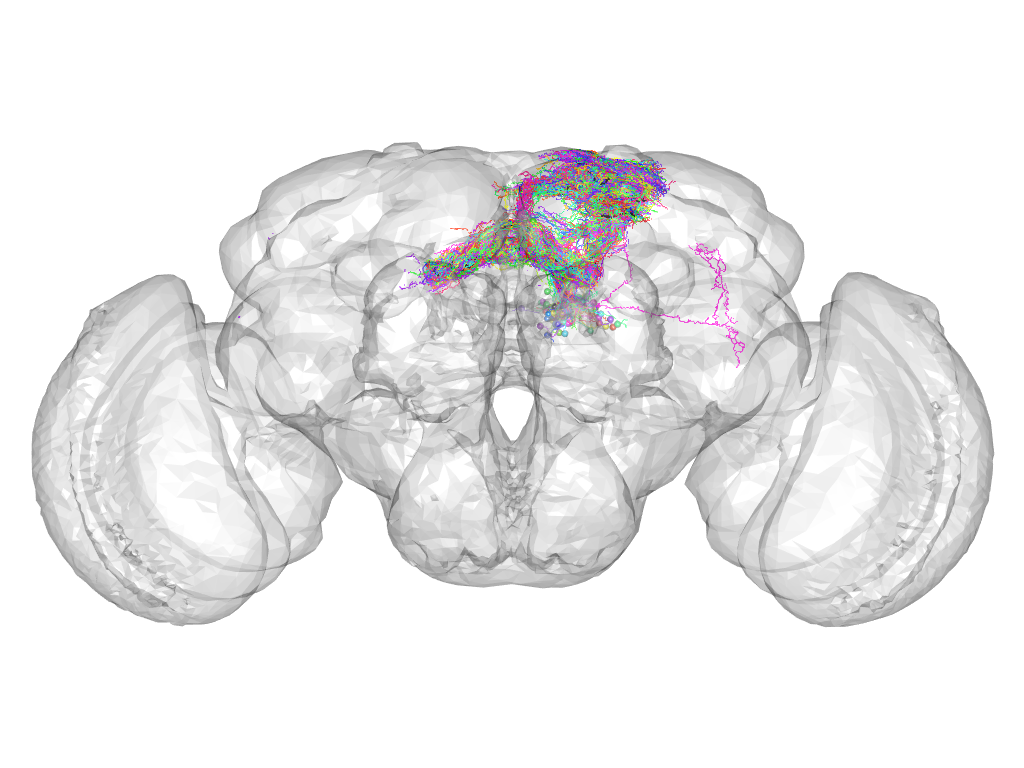
Cluster 141
Part of supercluster XIX
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (Trh-M-400106), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 887 | Trh-F-100046 | XIX | 0.58 |
| 83 | TH-M-300024 | XIX | 0.56 |
| 753 | VGlut-F-600178 | XIV | 0.40 |
| 189 | TH-M-200006 | XIX | 0.35 |
| 193 | TH-M-100039 | XIX | 0.29 |
Cluster composition
This cluster has 84 neurons. The exemplar of this cluster (Trh-M-400106) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| Trh-M-400106 | 2068 | TPHMARCM-M001571_seg001 | Trh-Gal4 | M | 27835.28 | 1.00 |
| Trh-M-400109 | 2071 | TPHMARCM-M001574_seg001 | Trh-Gal4 | M | 20526.03 | 0.73 |
| Trh-M-500195 | 2115 | TPHMARCM-M001613_seg001 | Trh-Gal4 | M | 18943.14 | 0.70 |
| Trh-M-200091 | 2123 | TPHMARCM-M001617_seg003 | Trh-Gal4 | M | 18406.30 | 0.67 |
| Trh-M-400123 | 2131 | TPHMARCM-M001622_seg001 | Trh-Gal4 | M | 19583.39 | 0.71 |
| Trh-M-400134 | 2151 | TPHMARCM-M001639_seg001 | Trh-Gal4 | M | 18283.48 | 0.66 |
| Trh-M-400136 | 2153 | TPHMARCM-M001641_seg001 | Trh-Gal4 | M | 20047.87 | 0.73 |
| Trh-M-200102 | 2202 | TPHMARCM-M001692_seg001 | Trh-Gal4 | M | 19317.04 | 0.72 |
| Trh-M-200114 | 2253 | TPHMARCM-M001740_seg001 | Trh-Gal4 | M | 19832.17 | 0.70 |
| Trh-M-200119 | 2269 | TPHMARCM-M001757_seg001 | Trh-Gal4 | M | 19260.46 | 0.71 |
| Trh-M-300139 | 2566 | TPHMARCM-M001543_seg001 | Trh-Gal4 | M | 19757.18 | 0.70 |
| Trh-M-200039 | 2581 | TPHMARCM-861M_seg1 | Trh-Gal4 | M | 20028.50 | 0.71 |
| Trh-M-700070 | 2592 | TPHMARCM-1176M_seg1 | Trh-Gal4 | M | 20195.78 | 0.73 |
| Trh-M-300128 | 2603 | TPHMARCM-1267M_seg1 | Trh-Gal4 | M | 18598.29 | 0.69 |
| Trh-M-200069 | 2643 | TPHMARCM-M001412_seg001 | Trh-Gal4 | M | 19757.00 | 0.72 |
| Trh-M-200071 | 2645 | TPHMARCM-M001414_seg001 | Trh-Gal4 | M | 19795.21 | 0.71 |
| Trh-M-300133 | 2725 | TPHMARCM-M001489_seg001 | Trh-Gal4 | M | 20323.44 | 0.74 |
| Trh-M-700086 | 2749 | TPHMARCM-M001515_seg001 | Trh-Gal4 | M | 19989.97 | 0.71 |
| Trh-M-400055 | 2811 | TPHMARCM-1040M_seg2 | Trh-Gal4 | M | 19980.77 | 0.73 |
| Trh-M-300093 | 2849 | TPHMARCM-1077M_seg1 | Trh-Gal4 | M | 19825.69 | 0.71 |
| Trh-M-300103 | 2864 | TPHMARCM-1102M_seg1 | Trh-Gal4 | M | 18622.79 | 0.69 |
| Trh-M-300105 | 2866 | TPHMARCM-1104M_seg1 | Trh-Gal4 | M | 19763.84 | 0.71 |
| Trh-M-100062 | 2895 | TPHMARCM-1134M_seg1 | Trh-Gal4 | M | 20203.08 | 0.73 |
| Trh-M-300115 | 2904 | TPHMARCM-1144M_seg1 | Trh-Gal4 | M | 19998.13 | 0.72 |
| Trh-M-200057 | 2918 | TPHMARCM-1169M_seg1 | Trh-Gal4 | M | 19689.96 | 0.72 |
| Trh-M-300075 | 2920 | TPHMARCM-853M_seg1 | Trh-Gal4 | M | 20562.56 | 0.73 |
| Trh-M-200037 | 2924 | TPHMARCM-860M_seg1 | Trh-Gal4 | M | 19541.29 | 0.71 |
| Trh-M-200042 | 2932 | TPHMARCM-867M_seg1 | Trh-Gal4 | M | 20174.93 | 0.72 |
| Trh-M-300085 | 2970 | TPHMARCM-910M_seg1 | Trh-Gal4 | M | 17037.61 | 0.63 |
| Trh-M-200044 | 2985 | TPHMARCM-922M_seg1 | Trh-Gal4 | M | 20185.77 | 0.72 |
| Trh-M-300089 | 3007 | TPHMARCM-956M_seg1 | Trh-Gal4 | M | 18064.05 | 0.65 |
| Trh-M-200007 | 3425 | TPHMARCM-118M_seg2 | Trh-Gal4 | M | 19818.00 | 0.71 |
| Trh-M-200009 | 3429 | TPHMARCM-164M_seg2 | Trh-Gal4 | M | 19399.15 | 0.72 |
| Trh-M-200018 | 3460 | TPHMARCM-321M_seg1 | Trh-Gal4 | M | 19449.86 | 0.71 |
| Trh-M-300008 | 3465 | TPHMARCM-335M_seg2 | Trh-Gal4 | M | 17393.26 | 0.66 |
| Trh-M-300009 | 3466 | TPHMARCM-336M_seg1 | Trh-Gal4 | M | 19004.53 | 0.71 |
| Trh-M-300012 | 3496 | TPHMARCM-366M_seg1 | Trh-Gal4 | M | 18168.59 | 0.65 |
| Trh-M-300040 | 3539 | TPHMARCM-445M_seg1 | Trh-Gal4 | M | 20368.65 | 0.73 |
| Trh-M-300049 | 3567 | TPHMARCM-498M_seg1 | Trh-Gal4 | M | 19374.00 | 0.70 |
| Trh-M-300054 | 3604 | TPHMARCM-547M_seg1 | Trh-Gal4 | M | 20275.89 | 0.73 |
| Trh-M-300057 | 3618 | TPHMARCM-582M_seg1 | Trh-Gal4 | M | 19163.07 | 0.69 |
| Trh-M-300060 | 3634 | TPHMARCM-616M_seg1 | Trh-Gal4 | M | 19469.76 | 0.71 |
| Trh-M-400030 | 3639 | TPHMARCM-629M_seg1 | Trh-Gal4 | M | 20519.89 | 0.73 |
| Trh-M-300066 | 3667 | TPHMARCM-706M_seg1 | Trh-Gal4 | M | 19910.50 | 0.73 |
| Trh-M-400037 | 3668 | TPHMARCM-710M_seg1 | Trh-Gal4 | M | 20681.91 | 0.74 |
| Trh-M-300068 | 3684 | TPHMARCM-798M_seg1 | Trh-Gal4 | M | 21125.97 | 0.76 |
| Trh-F-400025 | 10436 | TPHMARCM-172F_seg1 | Trh-Gal4 | F | 19592.39 | 0.70 |
| Trh-F-400089 | 10446 | TPHMARCM-1086F_seg2 | Trh-Gal4 | F | 18910.11 | 0.70 |
| Trh-F-200113 | 12380 | TPHMARCM-1175F_seg1 | Trh-Gal4 | F | 20466.03 | 0.73 |
| Trh-F-700056 | 12394 | TPHMARCM-1192F_seg1 | Trh-Gal4 | F | 20109.31 | 0.72 |
| Trh-F-200120 | 12414 | TPHMARCM-1207F_seg1 | Trh-Gal4 | F | 19217.11 | 0.69 |
| Trh-F-700061 | 12426 | TPHMARCM-1221F_seg1 | Trh-Gal4 | F | 20121.75 | 0.71 |
| Trh-F-600091 | 12443 | TPHMARCM-1237F_seg1 | Trh-Gal4 | F | 19024.70 | 0.69 |
| Trh-F-300144 | 12454 | TPHMARCM-1246F_seg1 | Trh-Gal4 | F | 19384.04 | 0.72 |
| Trh-F-100075 | 12515 | TPHMARCM-1297F_seg1 | Trh-Gal4 | F | 19071.55 | 0.70 |
| Trh-F-500185 | 12543 | TPHMARCM-1331F_seg1 | Trh-Gal4 | F | 20394.17 | 0.71 |
| Trh-F-200127 | 12550 | TPHMARCM-1337F_seg1 | Trh-Gal4 | F | 20042.12 | 0.72 |
| Trh-F-200128 | 12551 | TPHMARCM-1338F_seg1 | Trh-Gal4 | F | 16884.69 | 0.65 |
| Trh-F-300135 | 12702 | TPHMARCM-1151F_seg1 | Trh-Gal4 | F | 19325.14 | 0.72 |
| Trh-F-300138 | 12706 | TPHMARCM-1163F_seg1 | Trh-Gal4 | F | 19867.78 | 0.70 |
| Trh-F-200096 | 12712 | TPHMARCM-855F_seg1 | Trh-Gal4 | F | 20374.50 | 0.71 |
| Trh-F-300103 | 13219 | TPHMARCM-701F_seg1 | Trh-Gal4 | F | 20600.01 | 0.72 |
| Trh-F-500080 | 13221 | TPHMARCM-703F_seg1 | Trh-Gal4 | F | 20558.51 | 0.71 |
| Trh-F-400074 | 13231 | TPHMARCM-718F_seg1 | Trh-Gal4 | F | 19590.37 | 0.69 |
| Trh-F-200093 | 13302 | TPHMARCM-784F_seg1 | Trh-Gal4 | F | 20384.22 | 0.73 |
| Trh-F-100026 | 13832 | TPHMARCM-053F_seg1 | Trh-Gal4 | F | 18306.10 | 0.62 |
| Trh-F-200004 | 13869 | TPHMARCM-139F_seg1 | Trh-Gal4 | F | 19651.49 | 0.69 |
| Trh-F-200007 | 13912 | TPHMARCM-188F_seg1 | Trh-Gal4 | F | 20134.29 | 0.69 |
| Trh-F-200022 | 13947 | TPHMARCM-239F_seg1 | Trh-Gal4 | F | 19135.49 | 0.70 |
| Trh-F-300029 | 13999 | TPHMARCM-289F_seg1 | Trh-Gal4 | F | 20040.51 | 0.72 |
| Trh-F-300035 | 14011 | TPHMARCM-299F_seg1 | Trh-Gal4 | F | 19855.87 | 0.72 |
| Trh-F-300044 | 14060 | TPHMARCM-388F_seg1 | Trh-Gal4 | F | 19480.03 | 0.72 |
| Trh-F-300047 | 14073 | TPHMARCM-422F_seg1 | Trh-Gal4 | F | 20493.36 | 0.60 |
| Trh-F-400051 | 14074 | TPHMARCM-423F_seg1 | Trh-Gal4 | F | 20034.77 | 0.68 |
| Trh-F-300048 | 14075 | TPHMARCM-424F_seg1 | Trh-Gal4 | F | 20657.50 | 0.72 |
| Trh-F-300056 | 14083 | TPHMARCM-432F_seg1 | Trh-Gal4 | F | 19068.55 | 0.69 |
| Trh-F-200066 | 14093 | TPHMARCM-447F_seg1 | Trh-Gal4 | F | 20043.20 | 0.72 |
| Trh-F-500042 | 14111 | TPHMARCM-469F_seg1 | Trh-Gal4 | F | 20481.95 | 0.70 |
| Trh-F-300073 | 14139 | TPHMARCM-551F_seg1 | Trh-Gal4 | F | 19923.43 | 0.73 |
| Trh-F-500052 | 14170 | TPHMARCM-608F_seg1 | Trh-Gal4 | F | 19019.71 | 0.68 |
| Trh-F-400057 | 14175 | TPHMARCM-612F_seg1 | Trh-Gal4 | F | 20019.48 | 0.72 |
| Trh-F-300090 | 14186 | TPHMARCM-625F_seg1 | Trh-Gal4 | F | 19091.04 | 0.70 |
| Trh-F-300115 | 14285 | TPHMARCM-828F_seg1 | Trh-Gal4 | F | 20370.46 | 0.72 |
| VGlut-F-200285 | 15182 | DvGlutMARCM-F1513_seg1 | VGlut-Gal4 | F | 19219.21 | 0.69 |