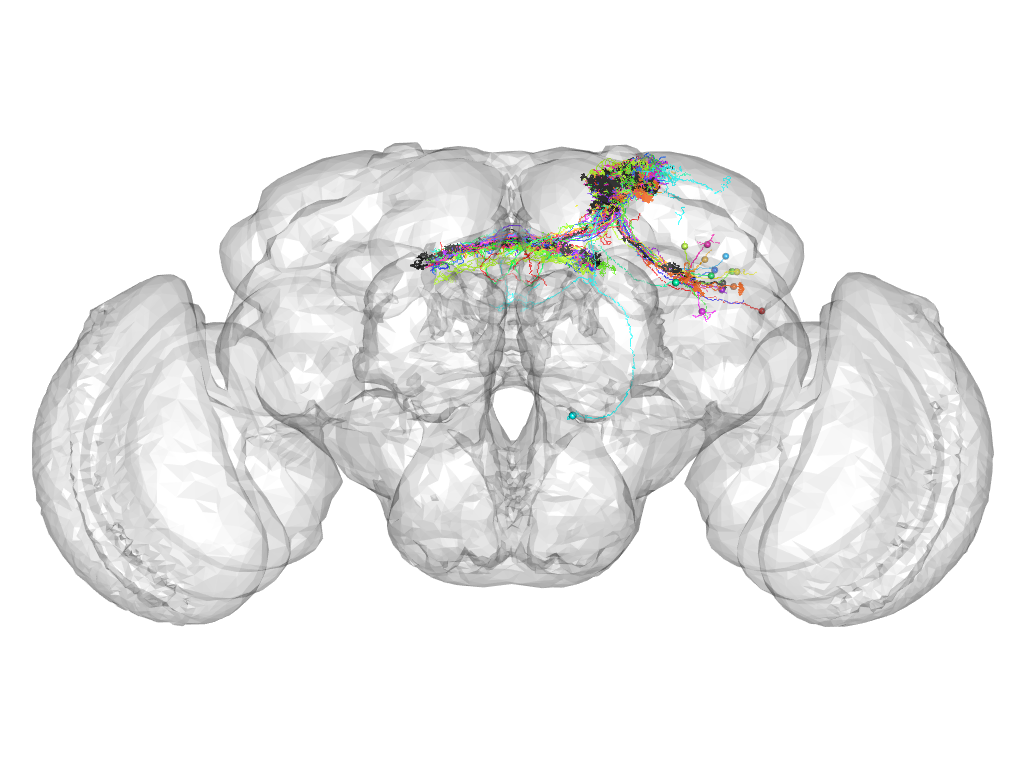
Cluster 189
Part of supercluster XIX
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (TH-M-200006), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 83 | TH-M-300024 | XIX | 0.71 |
| 886 | Trh-F-300042 | XIX | 0.47 |
| 193 | TH-M-100039 | XIX | 0.42 |
| 141 | Trh-M-400106 | XIX | 0.39 |
| 881 | Trh-F-200023 | XIX | 0.32 |
Cluster composition
This cluster has 19 neurons. The exemplar of this cluster (TH-M-200006) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| TH-M-200006 | 3166 | THMARCM-244M_seg1 | TH-Gal4 | M | 94542.00 | 1.00 |
| fru-M-500106 | 999 | FruMARCM-M001180_seg001 | fru-Gal4 | M | 52431.69 | 0.54 |
| TH-M-200009 | 3168 | THMARCM-256M_seg1 | TH-Gal4 | M | 70620.43 | 0.75 |
| TH-M-300032 | 3218 | THMARCM-348M_seg1 | TH-Gal4 | M | 69487.31 | 0.76 |
| TH-M-200092 | 3319 | THMARCM-567M_seg | TH-Gal4 | M | 71117.90 | 0.77 |
| TH-M-300059 | 3325 | THMARCM-579M_seg | TH-Gal4 | M | 54430.97 | 0.58 |
| Trh-M-200000 | 3413 | TPHMARCM-098M_seg1 | Trh-Gal4 | M | 62370.12 | 0.70 |
| Trh-M-300037 | 3534 | TPHMARCM-435M_seg1 | Trh-Gal4 | M | 58151.57 | 0.66 |
| Cha-F-300341 | 3917 | ChaMARCM-F001576_seg001 | Cha-Gal4 | F | 64816.31 | 0.68 |
| Cha-F-600039 | 7525 | ChaMARCM-F000376_seg001 | Cha-Gal4 | F | 9771.78 | 0.34 |
| Cha-F-500050 | 8119 | ChaMARCM-F000168_seg003 | Cha-Gal4 | F | 45547.15 | 0.37 |
| Gad1-F-300064 | 11227 | GadMARCM-F000209_seg002 | Gad1-Gal4 | F | 63851.22 | 0.73 |
| Trh-F-200109 | 12703 | TPHMARCM-1154F_seg1 | Trh-Gal4 | F | 63516.88 | 0.70 |
| TH-F-000008 | 13343 | THMARCM-096F_seg1 | TH-Gal4 | F | 68104.50 | 0.75 |
| TH-F-200054 | 13503 | THMARCM-378F_seg1 | TH-Gal4 | F | 69840.63 | 0.77 |
| TH-F-200066 | 13515 | THMARCM-391F_seg2 | TH-Gal4 | F | 70289.31 | 0.77 |
| TH-F-300065 | 13564 | THMARCM-466F_seg1 | TH-Gal4 | F | 71926.87 | 0.78 |
| TH-F-200098 | 13596 | THMARCM-529F_seg1 | TH-Gal4 | F | 71452.61 | 0.77 |
| Trh-F-300023 | 13982 | TPHMARCM-272F_seg1 | Trh-Gal4 | F | 63230.58 | 0.72 |