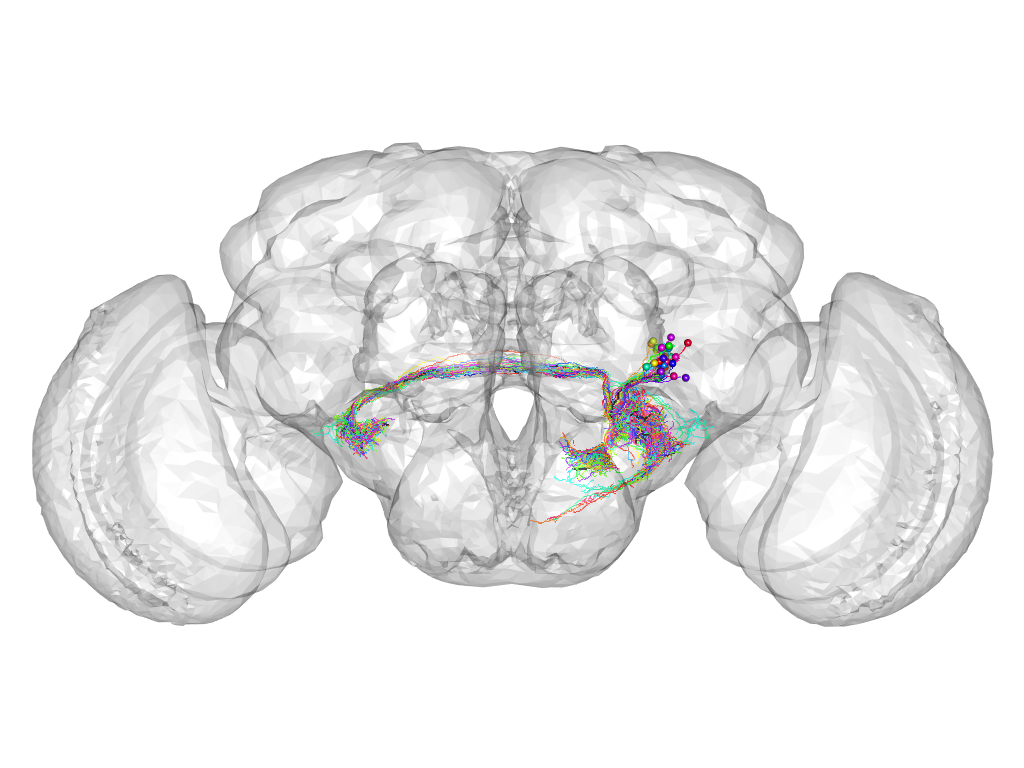
Cluster 803
Part of supercluster VIII
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (Trh-F-700063), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 801 | Trh-F-600102 | VIII | 0.65 |
| 804 | Trh-F-500188 | VIII | 0.55 |
| 960 | VGlut-F-500219 | VIII | 0.50 |
| 177 | Trh-M-300109 | VIII | 0.47 |
| 652 | Tdc2-F-200010 | VIII | 0.44 |
Cluster composition
This cluster has 32 neurons. The exemplar of this cluster (Trh-F-700063) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| Trh-F-700063 | 12482 | TPHMARCM-1271F_seg1 | Trh-Gal4 | F | 12368.70 | 1.00 |
| fru-M-600101 | 4 | FruMARCM-M002287_seg001 | fru-Gal4 | M | 8808.10 | 0.69 |
| fru-M-900032 | 1178 | FruMARCM-M001361_seg001 | fru-Gal4 | M | 8353.97 | 0.68 |
| fru-M-400110 | 1346 | FruMARCM-M001096_seg003 | fru-Gal4 | M | 8557.26 | 0.66 |
| Trh-M-600058 | 1993 | TPHMARCM-1012M_seg1 | Trh-Gal4 | M | 9120.40 | 0.73 |
| Trh-M-600082 | 2043 | TPHMARCM-M001551_seg003 | Trh-Gal4 | M | 8991.00 | 0.73 |
| Trh-M-700011 | 2928 | TPHMARCM-863M_seg1 | Trh-Gal4 | M | 8772.65 | 0.70 |
| Trh-M-300001 | 3417 | TPHMARCM-102M_seg1 | Trh-Gal4 | M | 9229.79 | 0.73 |
| Trh-M-500004 | 3436 | TPHMARCM-183M_seg1 | Trh-Gal4 | M | 9102.63 | 0.73 |
| Trh-M-500006 | 3442 | TPHMARCM-191M_seg1 | Trh-Gal4 | M | 8960.26 | 0.71 |
| Trh-M-600004 | 3545 | TPHMARCM-456M_seg1 | Trh-Gal4 | M | 8926.12 | 0.72 |
| Trh-M-500042 | 3646 | TPHMARCM-660M_seg2 | Trh-Gal4 | M | 9148.06 | 0.73 |
| Trh-M-400036 | 3649 | TPHMARCM-663M_seg1 | Trh-Gal4 | M | 9218.66 | 0.73 |
| Trh-M-500051 | 3671 | TPHMARCM-773M_seg1 | Trh-Gal4 | M | 9372.96 | 0.75 |
| VGlut-F-600728 | 6842 | DvGlutMARCM-F004022_seg002 | VGlut-Gal4 | F | 6978.91 | 0.54 |
| fru-F-200070 | 7384 | FruMARCM-F001027_seg002 | fru-Gal4 | F | 8505.48 | 0.72 |
| Cha-F-500086 | 8390 | ChaMARCM-F000327_seg001 | Cha-Gal4 | F | 7622.58 | 0.57 |
| VGlut-F-600589 | 9288 | DvGlutMARCM-F003336_seg001 | VGlut-Gal4 | F | 9047.96 | 0.73 |
| VGlut-F-300465 | 9319 | DvGlutMARCM-F003157_seg001 | VGlut-Gal4 | F | 8961.96 | 0.71 |
| fru-F-500008 | 9761 | FruMARCM-F000272_seg001 | fru-Gal4 | F | 9277.69 | 0.75 |
| Trh-F-500106 | 12382 | TPHMARCM-1178F_seg1 | Trh-Gal4 | F | 9317.16 | 0.76 |
| Trh-F-600071 | 12397 | TPHMARCM-1195F_seg1 | Trh-Gal4 | F | 9169.09 | 0.75 |
| Trh-F-600081 | 12422 | TPHMARCM-1213F_seg1 | Trh-Gal4 | F | 8769.48 | 0.71 |
| Trh-F-500148 | 12485 | TPHMARCM-1274F_seg1 | Trh-Gal4 | F | 9016.37 | 0.74 |
| Trh-F-500154 | 12498 | TPHMARCM-1283F_seg1 | Trh-Gal4 | F | 9419.09 | 0.74 |
| Trh-F-700018 | 13235 | TPHMARCM-722F_seg1 | Trh-Gal4 | F | 9142.86 | 0.74 |
| Trh-F-600054 | 13301 | TPHMARCM-783F_seg1 | Trh-Gal4 | F | 8951.23 | 0.72 |
| Trh-F-500093 | 13310 | TPHMARCM-800F_seg1 | Trh-Gal4 | F | 9198.93 | 0.75 |
| Trh-F-500039 | 14089 | TPHMARCM-440F_seg1 | Trh-Gal4 | F | 8964.44 | 0.73 |
| Trh-F-500050 | 14155 | TPHMARCM-579F_seg1 | Trh-Gal4 | F | 9478.86 | 0.77 |
| Trh-F-500064 | 14215 | TPHMARCM-654F_seg1 | Trh-Gal4 | F | 8795.25 | 0.67 |
| Trh-F-500073 | 14236 | TPHMARCM-691F_seg1 | Trh-Gal4 | F | 8731.36 | 0.72 |