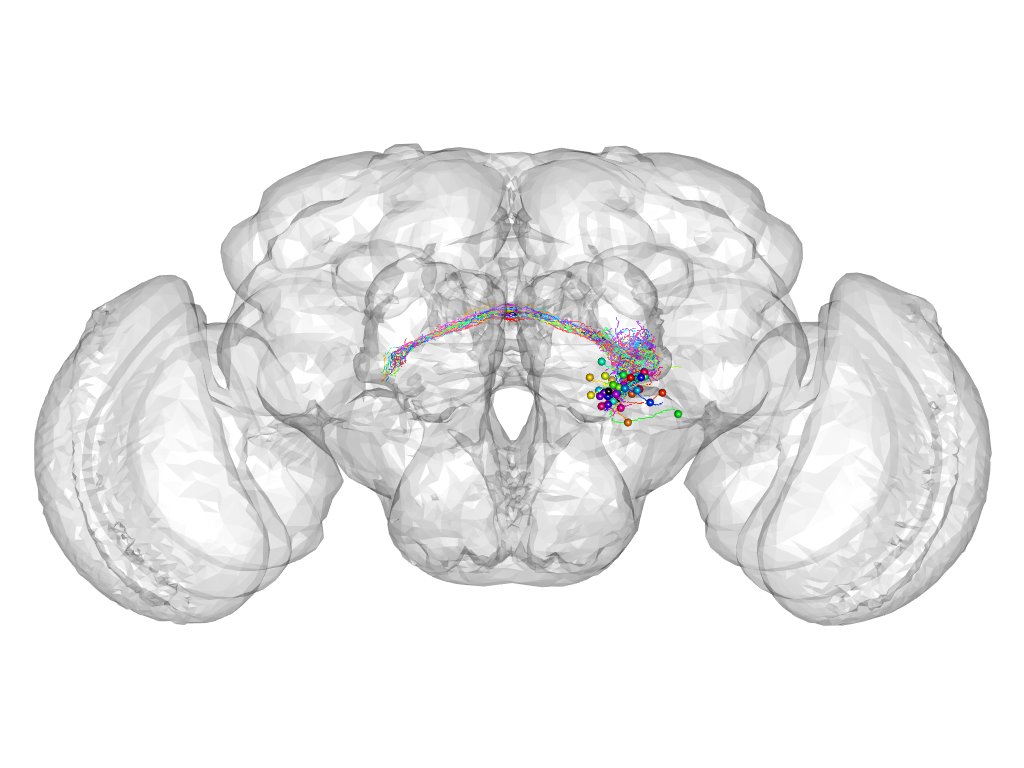
Cluster 555
Part of supercluster XII
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (VGlut-F-900099), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 997 | VGlut-F-400032 | XII | 0.78 |
| 420 | VGlut-F-700517 | XII | 0.69 |
| 653 | Trh-F-000043 | XIX | 0.62 |
| 545 | VGlut-F-700278 | XII | 0.55 |
| 662 | Tdc2-F-400026 | XIII | 0.54 |
| 835 | VGlut-F-400504 | XII | 0.53 |
Cluster composition
This cluster has 39 neurons. The exemplar of this cluster (VGlut-F-900099) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| VGlut-F-900099 | 8637 | DvGlutMARCM-F003689_seg002 | VGlut-Gal4 | F | 4760.70 | 1.00 |
| fru-M-500084 | 1437 | FruMARCM-M001009_seg001 | fru-Gal4 | M | 3306.26 | 0.67 |
| fru-M-100057 | 1588 | FruMARCM-M000655_seg001 | fru-Gal4 | M | 3866.59 | 0.78 |
| fru-M-200084 | 1617 | FruMARCM-M000729_seg002 | fru-Gal4 | M | 3460.43 | 0.71 |
| fru-M-300039 | 1785 | FruMARCM-M000313_seg001 | fru-Gal4 | M | 3751.88 | 0.75 |
| VGlut-F-800287 | 5694 | DvGlutMARCM-F004413_seg001 | VGlut-Gal4 | F | 3382.16 | 0.73 |
| VGlut-F-800292 | 5734 | DvGlutMARCM-F004440_seg002 | VGlut-Gal4 | F | 3748.48 | 0.75 |
| VGlut-F-700561 | 5742 | DvGlutMARCM-F004449_seg001 | VGlut-Gal4 | F | 4031.94 | 0.81 |
| VGlut-F-500850 | 5762 | DvGlutMARCM-F004465_seg001 | VGlut-Gal4 | F | 3810.89 | 0.79 |
| VGlut-F-400920 | 5830 | DvGlutMARCM-F004519_seg001 | VGlut-Gal4 | F | 3862.01 | 0.80 |
| VGlut-F-800324 | 5931 | DvGlutMARCM-F004591_seg001 | VGlut-Gal4 | F | 3855.36 | 0.77 |
| VGlut-F-900128 | 6507 | DvGlutMARCM-F004285_seg002 | VGlut-Gal4 | F | 3916.02 | 0.81 |
| VGlut-F-500827 | 6615 | DvGlutMARCM-F004368_seg001 | VGlut-Gal4 | F | 3806.66 | 0.76 |
| VGlut-F-500833 | 6623 | DvGlutMARCM-F004376_seg001 | VGlut-Gal4 | F | 3907.94 | 0.80 |
| VGlut-F-900130 | 6641 | DvGlutMARCM-F004388_seg002 | VGlut-Gal4 | F | 3699.41 | 0.74 |
| VGlut-F-600788 | 6652 | DvGlutMARCM-F004395_seg001 | VGlut-Gal4 | F | 3686.39 | 0.78 |
| VGlut-F-700456 | 6755 | DvGlutMARCM-F003959_seg001 | VGlut-Gal4 | F | 3387.00 | 0.74 |
| VGlut-F-600741 | 6882 | DvGlutMARCM-F004040_seg001 | VGlut-Gal4 | F | 3915.43 | 0.80 |
| VGlut-F-700475 | 6922 | DvGlutMARCM-F004076_seg001 | VGlut-Gal4 | F | 3609.17 | 0.75 |
| VGlut-F-700272 | 8508 | DvGlutMARCM-F003091_seg001 | VGlut-Gal4 | F | 3873.14 | 0.78 |
| VGlut-F-500575 | 8561 | DvGlutMARCM-F003128_seg001 | VGlut-Gal4 | F | 3668.46 | 0.77 |
| VGlut-F-500723 | 8598 | DvGlutMARCM-F003661_seg001 | VGlut-Gal4 | F | 3721.20 | 0.77 |
| VGlut-F-500685 | 8948 | DvGlutMARCM-F003513_seg001 | VGlut-Gal4 | F | 3646.91 | 0.74 |
| VGlut-F-500694 | 8993 | DvGlutMARCM-F003542_seg001 | VGlut-Gal4 | F | 3780.11 | 0.77 |
| VGlut-F-700332 | 9039 | DvGlutMARCM-F003362_seg001 | VGlut-Gal4 | F | 3793.29 | 0.78 |
| VGlut-F-600600 | 9047 | DvGlutMARCM-F003369_seg001 | VGlut-Gal4 | F | 3654.54 | 0.72 |
| VGlut-F-600612 | 9140 | DvGlutMARCM-F003431_seg002 | VGlut-Gal4 | F | 3709.84 | 0.78 |
| VGlut-F-500604 | 9385 | DvGlutMARCM-F003214_seg002 | VGlut-Gal4 | F | 3676.90 | 0.77 |
| VGlut-F-200170 | 10261 | DvGlutMARCM-F914_seg1 | VGlut-Gal4 | F | 3766.98 | 0.76 |
| VGlut-F-500487 | 11014 | DvGlutMARCM-F002898_seg003 | VGlut-Gal4 | F | 3748.20 | 0.72 |
| VGlut-F-800096 | 11065 | DvGlutMARCM-F002943_seg001 | VGlut-Gal4 | F | 3833.96 | 0.77 |
| VGlut-F-600199 | 11761 | DvGlutMARCM-F002332_seg002 | VGlut-Gal4 | F | 3543.16 | 0.73 |
| VGlut-F-700061 | 11872 | DvGlutMARCM-F002417_seg001 | VGlut-Gal4 | F | 3848.83 | 0.79 |
| VGlut-F-500291 | 12821 | DvGlutMARCM-F1869_seg1 | VGlut-Gal4 | F | 3718.74 | 0.75 |
| VGlut-F-600019 | 14441 | DvGlutMARCM-F732_seg1 | VGlut-Gal4 | F | 3863.37 | 0.79 |
| VGlut-F-400207 | 14498 | DvGlutMARCM-F769_seg1 | VGlut-Gal4 | F | 3794.56 | 0.75 |
| VGlut-F-400276 | 14665 | DvGlutMARCM-F926_seg1 | VGlut-Gal4 | F | 3654.28 | 0.75 |
| VGlut-F-000386 | 15281 | DvGlutMARCM-F1607_seg1 | VGlut-Gal4 | F | 3589.28 | 0.74 |
| VGlut-F-300311 | 15300 | DvGlutMARCM-F1634_seg1 | VGlut-Gal4 | F | 3737.25 | 0.77 |