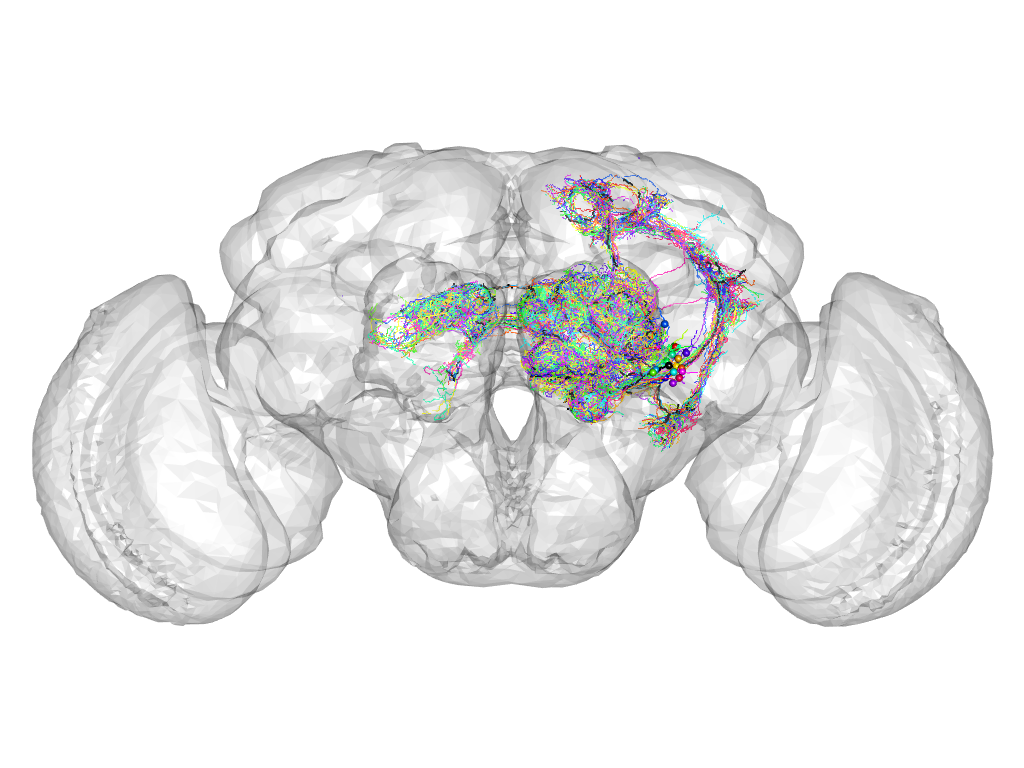
Cluster 653
Part of supercluster XIX
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (Trh-F-000043), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 662 | Tdc2-F-400026 | XIII | 0.47 |
| 994 | VGlut-F-200045 | XIX | 0.35 |
| 806 | Trh-F-100085 | XII | 0.30 |
| 144 | Trh-M-400113 | XII | 0.24 |
| 201 | Trh-M-100007 | XII | 0.21 |
Cluster composition
This cluster has 19 neurons. The exemplar of this cluster (Trh-F-000043) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| Trh-F-000043 | 10204 | TPHMARCM-1266F_seg1 | Trh-Gal4 | F | 61274.06 | 1.00 |
| Trh-M-000070 | 2231 | TPHMARCM-M001718_seg001 | Trh-Gal4 | M | 44503.14 | 0.72 |
| Trh-M-000039 | 2684 | TPHMARCM-M001451_seg001 | Trh-Gal4 | M | 44226.10 | 0.73 |
| Trh-M-000041 | 2686 | TPHMARCM-M001453_seg001 | Trh-Gal4 | M | 44851.04 | 0.72 |
| Trh-M-000008 | 3404 | TPHMARCM-090M_seg1 | Trh-Gal4 | M | 46406.83 | 0.73 |
| Trh-M-100037 | 3536 | TPHMARCM-439M_seg1 | Trh-Gal4 | M | 43769.46 | 0.71 |
| VGlut-F-000651 | 6463 | DvGlutMARCM-F004248_seg001 | VGlut-Gal4 | F | 25966.27 | 0.57 |
| VGlut-F-000624 | 6891 | DvGlutMARCM-F004047_seg001 | VGlut-Gal4 | F | 44818.39 | 0.72 |
| Trh-F-000009 | 10442 | TPHMARCM-039F_seg1 | Trh-Gal4 | F | 44778.79 | 0.72 |
| VGlut-F-100003 | 10763 | DvGlutMARCM-F-0303-03_F073_seg1 | VGlut-Gal4 | F | 45389.31 | 0.71 |
| VGlut-F-100272 | 11640 | DvGlutMARCM-F002215-VnewoACT_seg1 | VGlut-Gal4 | F | 21208.97 | 0.50 |
| Trh-F-000041 | 12431 | TPHMARCM-1227F_seg1 | Trh-Gal4 | F | 53238.59 | 0.88 |
| VGlut-F-000430 | 12965 | DvGlutMARCM-F1996_seg1 | VGlut-Gal4 | F | 44045.28 | 0.71 |
| Trh-F-000008 | 13816 | TPHMARCM-035F_seg1 | Trh-Gal4 | F | 45104.09 | 0.72 |
| Trh-F-100032 | 13842 | TPHMARCM-110F_seg1 | Trh-Gal4 | F | 44420.59 | 0.72 |
| Trh-F-100037 | 14019 | TPHMARCM-306F_seg1 | Trh-Gal4 | F | 43383.30 | 0.71 |
| Trh-F-100038 | 14023 | TPHMARCM-310F_seg1 | Trh-Gal4 | F | 44682.14 | 0.72 |
| VGlut-F-000189 | 14301 | DVGlutMARCM-F-0509-02-montage_seg1 | VGlut-Gal4 | F | 31960.69 | 0.57 |
| VGlut-F-100184 | 15370 | DvGlutMARCM-F1721_seg1 | VGlut-Gal4 | F | 16464.22 | 0.46 |