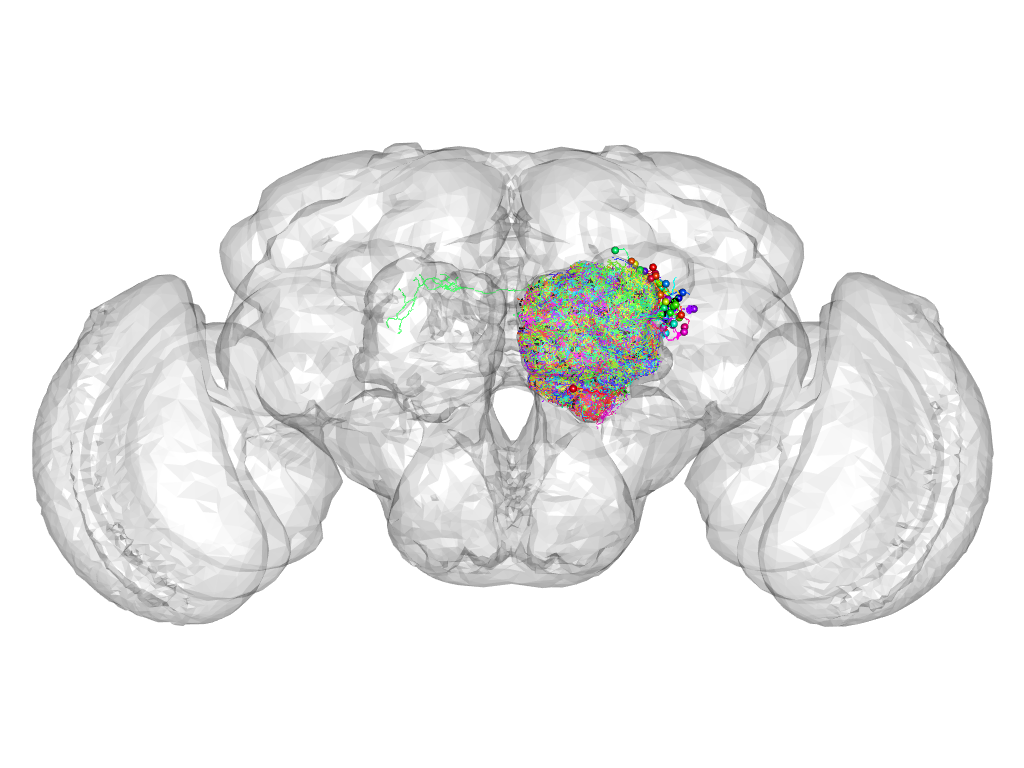
Cluster 144
Part of supercluster XII
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (Trh-M-400113), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 806 | Trh-F-100085 | XII | 0.79 |
| 662 | Tdc2-F-400026 | XIII | 0.71 |
| 201 | Trh-M-100007 | XII | 0.66 |
| 653 | Trh-F-000043 | XIX | 0.61 |
| 400 | VGlut-F-500072 | XII | 0.57 |
| 172 | Trh-M-100092 | XII | 0.56 |
| 774 | VGlut-F-500418 | XII | 0.51 |
Cluster composition
This cluster has 56 neurons. The exemplar of this cluster (Trh-M-400113) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| Trh-M-400113 | 2075 | TPHMARCM-M001577_seg002 | Trh-Gal4 | M | 86341.75 | 1.00 |
| Trh-M-300114 | 1995 | TPHMARCM-1143M_seg001 | Trh-Gal4 | M | 62497.12 | 0.74 |
| Trh-M-200076 | 2077 | TPHMARCM-M001579_seg001 | Trh-Gal4 | M | 62003.63 | 0.74 |
| Trh-M-300147 | 2157 | TPHMARCM-M001646_seg001 | Trh-Gal4 | M | 64349.86 | 0.74 |
| Trh-M-400137 | 2162 | TPHMARCM-M001651_seg001 | Trh-Gal4 | M | 58565.02 | 0.72 |
| Trh-M-300155 | 2171 | TPHMARCM-M001661_seg001 | Trh-Gal4 | M | 64305.24 | 0.75 |
| Trh-M-200127 | 2279 | TPHMARCM-M001762_seg001 | Trh-Gal4 | M | 57331.29 | 0.71 |
| 5-HT1B-M-500002 | 2489 | 5HT1bMARCM-M000035_seg001 | 5-HT1B-Gal4 | M | 64743.82 | 0.76 |
| 5-HT1B-M-400003 | 2499 | 5HT1bMARCM-M000044_seg002 | 5-HT1B-Gal4 | M | 61677.63 | 0.74 |
| 5-HT1B-M-400009 | 2509 | 5HT1bMARCM-M000052_seg001 | 5-HT1B-Gal4 | M | 32268.87 | 0.57 |
| 5-HT1B-M-300000 | 2510 | 5HT1bMARCM-M000053_seg001 | 5-HT1B-Gal4 | M | 61526.61 | 0.73 |
| 5-HT1B-M-400010 | 2514 | 5HT1bMARCM-M000056_seg001 | 5-HT1B-Gal4 | M | 61722.46 | 0.74 |
| 5-HT1B-M-400015 | 2523 | 5HT1bMARCM-M000064_seg001 | 5-HT1B-Gal4 | M | 47979.17 | 0.66 |
| 5-HT1B-M-400017 | 2527 | 5HT1bMARCM-M000068_seg001 | 5-HT1B-Gal4 | M | 63932.94 | 0.75 |
| Trh-M-300141 | 2568 | TPHMARCM-M001544_seg002 | Trh-Gal4 | M | 64228.95 | 0.75 |
| Trh-M-200059 | 2591 | TPHMARCM-1174M_seg1 | Trh-Gal4 | M | 64339.14 | 0.75 |
| Trh-M-300124 | 2599 | TPHMARCM-1218M_seg1 | Trh-Gal4 | M | 60650.21 | 0.72 |
| Trh-M-600068 | 2611 | TPHMARCM-1317M_seg1 | Trh-Gal4 | M | 58642.57 | 0.72 |
| Trh-M-500102 | 2681 | TPHMARCM-M001447_seg001 | Trh-Gal4 | M | 65950.21 | 0.76 |
| Trh-M-100066 | 2899 | TPHMARCM-1138M_seg1 | Trh-Gal4 | M | 64991.67 | 0.76 |
| Trh-M-300074 | 2919 | TPHMARCM-852M_seg1 | Trh-Gal4 | M | 61539.17 | 0.74 |
| TH-M-200054 | 3235 | THMARCM-379M_seg1 | TH-Gal4 | M | 33978.23 | 0.42 |
| TH-M-200089 | 3298 | THMARCM-510M_seg1 | TH-Gal4 | M | 39420.35 | 0.58 |
| Trh-M-300014 | 3511 | TPHMARCM-392M_seg1 | Trh-Gal4 | M | 64791.15 | 0.76 |
| Trh-M-300015 | 3512 | TPHMARCM-393M_seg1 | Trh-Gal4 | M | 46218.64 | 0.66 |
| Trh-M-300016 | 3513 | TPHMARCM-394M_seg1 | Trh-Gal4 | M | 63116.70 | 0.74 |
| Trh-M-300021 | 3518 | TPHMARCM-398M_seg1 | Trh-Gal4 | M | 60612.48 | 0.73 |
| Trh-M-300044 | 3557 | TPHMARCM-487M_seg2 | Trh-Gal4 | M | 56277.46 | 0.71 |
| Trh-M-200028 | 3596 | TPHMARCM-539M_seg1 | Trh-Gal4 | M | 64974.31 | 0.75 |
| Gad1-F-300137 | 3992 | GadMARCM-F000612_seg001 | Gad1-Gal4 | F | 48239.95 | 0.66 |
| Cha-F-600118 | 5439 | ChaMARCM-F000781_seg001 | Cha-Gal4 | F | 45746.82 | 0.64 |
| VGlut-F-000560 | 8628 | DvGlutMARCM-F003681_seg001 | VGlut-Gal4 | F | 42829.71 | 0.62 |
| Gad1-F-600063 | 9649 | GadMARCM-F000502_seg001 | Gad1-Gal4 | F | 48766.56 | 0.66 |
| Gad1-F-300048 | 10697 | GadMARCM-F000196_seg001 | Gad1-Gal4 | F | 37370.33 | 0.59 |
| 5-HT1B-F-500019 | 11199 | 5HT1bMARCM-F000029_seg001 | 5-HT1B-Gal4 | F | 59321.38 | 0.73 |
| Gad1-F-900029 | 11301 | GadMARCM-F000274_seg002 | Gad1-Gal4 | F | 43388.73 | 0.62 |
| 5-HT1B-F-500003 | 11608 | 5HT1bMARCM-F000005_seg001 | 5-HT1B-Gal4 | F | 53488.22 | 0.69 |
| 5-HT1B-F-500005 | 11610 | 5HT1bMARCM-F000007_seg001 | 5-HT1B-Gal4 | F | 56507.72 | 0.71 |
| 5-HT1B-F-500006 | 11611 | 5HT1bMARCM-F000007_seg002 | 5-HT1B-Gal4 | F | 47868.15 | 0.65 |
| 5-HT1B-F-500008 | 11613 | 5HT1bMARCM-F000009_seg001 | 5-HT1B-Gal4 | F | 59488.05 | 0.72 |
| 5-HT1B-F-500009 | 11614 | 5HT1bMARCM-F000010_seg001 | 5-HT1B-Gal4 | F | 57397.79 | 0.72 |
| VGlut-F-600294 | 12025 | DvGlutMARCM-F002544_seg001 | VGlut-Gal4 | F | 57365.52 | 0.71 |
| Trh-F-600106 | 12496 | TPHMARCM-1281F_seg2 | Trh-Gal4 | F | 62090.77 | 0.74 |
| Trh-F-300145 | 12522 | TPHMARCM-1304F_seg1 | Trh-Gal4 | F | 64280.18 | 0.74 |
| Trh-F-500186 | 12544 | TPHMARCM-1332F_seg1 | Trh-Gal4 | F | 62450.95 | 0.75 |
| Trh-F-700065 | 12558 | TPHMARCM-1346F_seg1 | Trh-Gal4 | F | 57776.27 | 0.72 |
| Trh-F-600107 | 12561 | TPHMARCM-1349F_seg1 | Trh-Gal4 | F | 63888.92 | 0.75 |
| Trh-F-200104 | 12696 | TPHMARCM-1120F_seg1 | Trh-Gal4 | F | 52268.07 | 0.67 |
| TH-F-700020 | 13366 | THMARCM-110_seg1 | TH-Gal4 | F | 47120.69 | 0.64 |
| TH-F-700029 | 13375 | THMARCM-134F_seg1 | TH-Gal4 | F | 30848.35 | 0.55 |
| TH-F-700033 | 13379 | THMARCM-139F_seg1 | TH-Gal4 | F | 34549.91 | 0.54 |
| TH-F-200080 | 13539 | THMARCM-427F_seg1 | TH-Gal4 | F | 32556.34 | 0.54 |
| Trh-F-300050 | 14077 | TPHMARCM-426F_seg1 | Trh-Gal4 | F | 55870.73 | 0.70 |
| Trh-F-500040 | 14109 | TPHMARCM-468F_seg1 | Trh-Gal4 | F | 63029.23 | 0.74 |
| Trh-F-200068 | 14112 | TPHMARCM-470F_seg1 | Trh-Gal4 | F | 63349.76 | 0.75 |
| VGlut-F-500135 | 14648 | DvGlutMARCM-F902_seg1 | VGlut-Gal4 | F | 53878.07 | 0.68 |