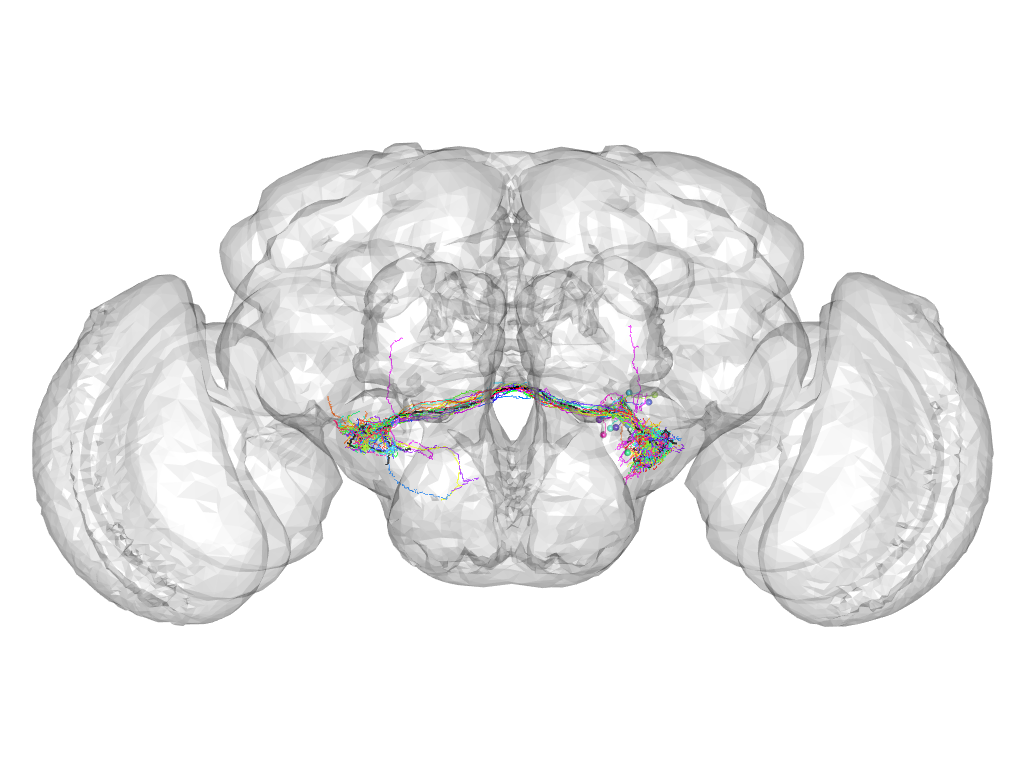
Cluster 459
Part of supercluster VIII
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (VGlut-F-500779), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 320 | Cha-F-400233 | VIII | 0.68 |
| 902 | VGlut-F-400300 | VIII | 0.67 |
| 309 | fru-F-700160 | VIII | 0.57 |
| 410 | fru-F-400147 | VIII | 0.57 |
| 27 | fru-M-300302 | VIII | 0.56 |
| 47 | fru-M-300198 | VIII | 0.56 |
Cluster composition
This cluster has 18 neurons. The exemplar of this cluster (VGlut-F-500779) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| VGlut-F-500779 | 7071 | DvGlutMARCM-F004200_seg001 | VGlut-Gal4 | F | 10671.71 | 1.00 |
| fru-M-800077 | 656 | FruMARCM-M001867_seg001 | fru-Gal4 | M | 7396.50 | 0.71 |
| fru-M-300148 | 1151 | FruMARCM-M001343_seg002 | fru-Gal4 | M | 7512.24 | 0.71 |
| fru-M-400137 | 1195 | FruMARCM-M001375_seg001 | fru-Gal4 | M | 7263.31 | 0.71 |
| fru-M-400150 | 1291 | FruMARCM-M001478_seg001 | fru-Gal4 | M | 7788.46 | 0.70 |
| fru-M-300062 | 1909 | FruMARCM-M000461_seg001 | fru-Gal4 | M | 7814.73 | 0.73 |
| fru-M-200056 | 1927 | FruMARCM-M000521_seg001 | fru-Gal4 | M | 6938.13 | 0.69 |
| VGlut-F-600790 | 5702 | DvGlutMARCM-F004414_seg001 | VGlut-Gal4 | F | 7651.20 | 0.74 |
| fru-F-700127 | 6251 | FruMARCM-F001265_seg001 | fru-Gal4 | F | 7740.42 | 0.75 |
| VGlut-F-700534 | 6529 | DvGlutMARCM-F004299_seg001 | VGlut-Gal4 | F | 7455.48 | 0.73 |
| VGlut-F-600701 | 6689 | DvGlutMARCM-F003908_seg001 | VGlut-Gal4 | F | 7444.19 | 0.71 |
| VGlut-F-700466 | 6771 | DvGlutMARCM-F003973_seg001 | VGlut-Gal4 | F | 7526.46 | 0.68 |
| VGlut-F-600705 | 6774 | DvGlutMARCM-F003974_seg001 | VGlut-Gal4 | F | 7477.63 | 0.73 |
| VGlut-F-400825 | 6832 | DvGlutMARCM-F004015_seg001 | VGlut-Gal4 | F | 7869.85 | 0.76 |
| VGlut-F-700418 | 7967 | DvGlutMARCM-F003794_seg004 | VGlut-Gal4 | F | 7206.18 | 0.64 |
| Cha-F-200061 | 8257 | ChaMARCM-F000243_seg001 | Cha-Gal4 | F | 5353.95 | 0.46 |
| fru-F-400009 | 11466 | FruMARCM-F000079_seg001 | fru-Gal4 | F | 7761.48 | 0.73 |
| VGlut-F-600197 | 11759 | DvGlutMARCM-F002331_seg001 | VGlut-Gal4 | F | 7095.89 | 0.70 |