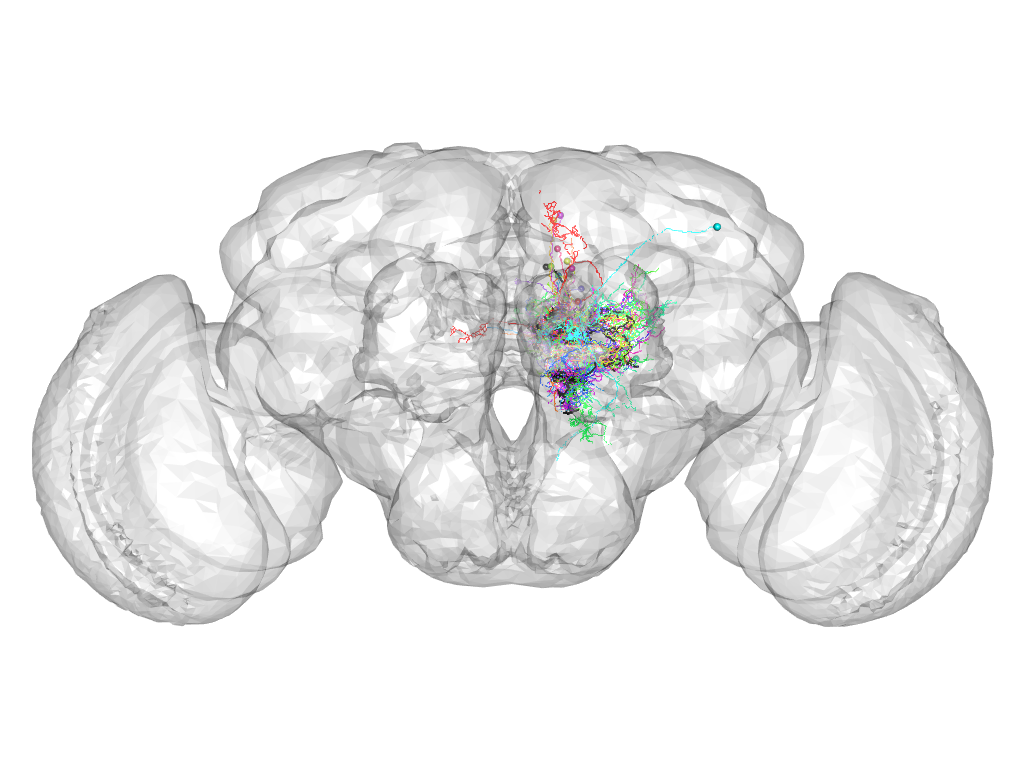
Cluster 380
Part of supercluster VI
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (VGlut-F-200603), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 666 | TH-F-200011 | IX | 0.58 |
| 707 | VGlut-F-200430 | VII | 0.54 |
| 425 | VGlut-F-500805 | VII | 0.54 |
| 872 | TH-F-100101 | IX | 0.52 |
| 826 | VGlut-F-500316 | VI | 0.51 |
| 353 | Cha-F-200264 | VI | 0.51 |
Cluster composition
This cluster has 17 neurons. The exemplar of this cluster (VGlut-F-200603) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| VGlut-F-200603 | 5879 | DvGlutMARCM-F004560_seg001 | VGlut-Gal4 | F | 29691.72 | 1.00 |
| Cha-F-500180 | 3840 | ChaMARCM-F001514_seg001 | Cha-Gal4 | F | 17225.78 | 0.53 |
| VGlut-F-400882 | 6426 | DvGlutMARCM-F004218_seg002 | VGlut-Gal4 | F | 15017.59 | 0.57 |
| VGlut-F-500806 | 6542 | DvGlutMARCM-F004307_seg001 | VGlut-Gal4 | F | 14825.40 | 0.60 |
| VGlut-F-700536 | 6569 | DvGlutMARCM-F004331_seg001 | VGlut-Gal4 | F | 13859.69 | 0.58 |
| VGlut-F-400829 | 6850 | DvGlutMARCM-F004027_seg001 | VGlut-Gal4 | F | 15455.96 | 0.61 |
| VGlut-F-300599 | 7074 | DvGlutMARCM-F004203_seg001 | VGlut-Gal4 | F | 21468.61 | 0.69 |
| Cha-F-800003 | 7463 | ChaMARCM-F000343_seg001 | Cha-Gal4 | F | 14312.60 | 0.49 |
| Cha-F-700115 | 7589 | ChaMARCM-F000442_seg003 | Cha-Gal4 | F | 15508.02 | 0.51 |
| Cha-F-200049 | 8239 | ChaMARCM-F000235_seg001 | Cha-Gal4 | F | 10544.68 | 0.40 |
| VGlut-F-200472 | 9218 | DvGlutMARCM-F003283_seg002 | VGlut-Gal4 | F | 17999.88 | 0.66 |
| Gad1-F-300116 | 9595 | GadMARCM-F000464_seg001 | Gad1-Gal4 | F | 16590.44 | 0.60 |
| VGlut-F-400610 | 10920 | DvGlutMARCM-F002817_seg001 | VGlut-Gal4 | F | 21383.27 | 0.72 |
| VGlut-F-700073 | 12079 | DvGlutMARCM-F002588_seg001 | VGlut-Gal4 | F | 15407.80 | 0.60 |
| VGlut-F-300136 | 14417 | DvGlutMARCM-F712_seg1 | VGlut-Gal4 | F | 20236.73 | 0.68 |
| VGlut-F-100157 | 15179 | DvGlutMARCM-F1509_seg1 | VGlut-Gal4 | F | 20857.05 | 0.67 |
| VGlut-F-500083 | 16053 | DvGlutMARCM-F440_seg4 | VGlut-Gal4 | F | 14086.89 | 0.60 |