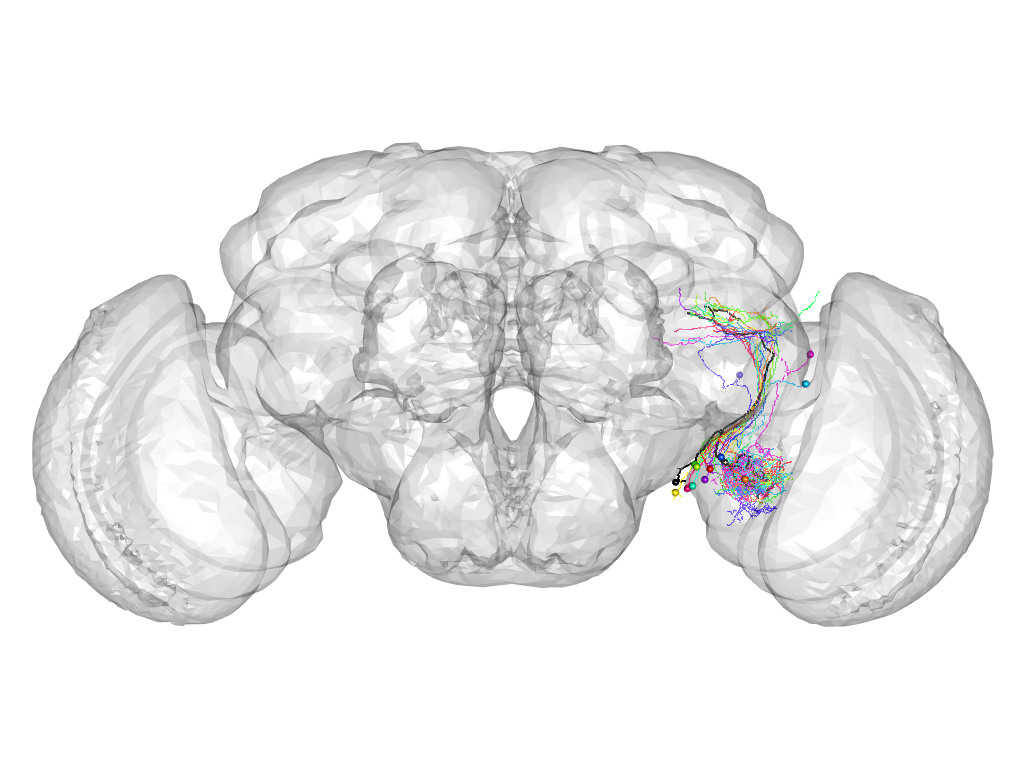
Cluster 369
Part of supercluster II
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (Cha-F-200187), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 775 | VGlut-F-500422 | II | 0.55 |
| 1022 | VGlut-F-400066 | II | 0.55 |
| 509 | VGlut-F-200536 | VII | 0.49 |
| 678 | Gad1-F-200035 | II | 0.39 |
| 263 | Cha-F-000316 | II | 0.35 |
Cluster composition
This cluster has 14 neurons. The exemplar of this cluster (Cha-F-200187) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| Cha-F-200187 | 5676 | ChaMARCM-F000938_seg001 | Cha-Gal4 | F | 12266.20 | 1.00 |
| fru-M-300217 | 677 | FruMARCM-M001883_seg001 | fru-Gal4 | M | 7186.86 | 0.56 |
| fru-M-700033 | 1889 | FruMARCM-M000424_seg002 | fru-Gal4 | M | 7000.37 | 0.60 |
| Cha-F-400189 | 5530 | ChaMARCM-F000838_seg001 | Cha-Gal4 | F | 6005.82 | 0.53 |
| VGlut-F-400911 | 5811 | DvGlutMARCM-F004506_seg003 | VGlut-Gal4 | F | 7964.78 | 0.65 |
| VGlut-F-500765 | 6826 | DvGlutMARCM-F004009_seg001 | VGlut-Gal4 | F | 6328.58 | 0.53 |
| VGlut-F-700510 | 7052 | DvGlutMARCM-F004181_seg001 | VGlut-Gal4 | F | 5433.60 | 0.40 |
| Cha-F-200073 | 7592 | ChaMARCM-F000444_seg001 | Cha-Gal4 | F | 8576.97 | 0.70 |
| Cha-F-700023 | 8041 | ChaMARCM-F000121_seg001 | Cha-Gal4 | F | 6639.58 | 0.53 |
| VGlut-F-700290 | 8527 | DvGlutMARCM-F003105_seg002 | VGlut-Gal4 | F | 6718.33 | 0.56 |
| Cha-F-300024 | 10112 | ChaMARCM-F000065_seg001 | Cha-Gal4 | F | 5424.35 | 0.38 |
| Gad1-F-800003 | 10601 | GadMARCM-F000112_seg002 | Gad1-Gal4 | F | 4797.61 | 0.46 |
| VGlut-F-100210 | 12920 | DvGlutMARCM-F1959_seg1 | VGlut-Gal4 | F | 6604.82 | 0.53 |
| Gad1-F-300003 | 13173 | GadMARCM-F010_seg2 | Gad1-Gal4 | F | 8031.80 | 0.64 |