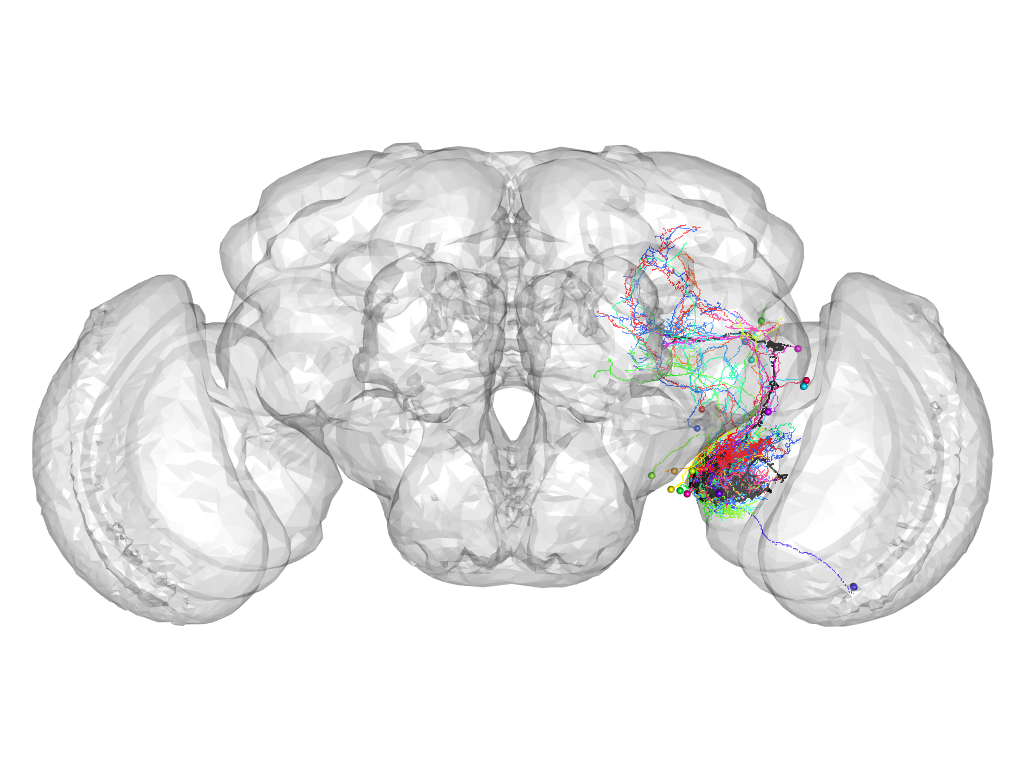
Cluster 1022
Part of supercluster II
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (VGlut-F-400066), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 775 | VGlut-F-500422 | II | 0.76 |
| 678 | Gad1-F-200035 | II | 0.62 |
| 509 | VGlut-F-200536 | VII | 0.61 |
| 228 | Cha-F-500191 | II | 0.44 |
| 640 | fru-F-500062 | II | 0.44 |
Cluster composition
This cluster has 20 neurons. The exemplar of this cluster (VGlut-F-400066) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| VGlut-F-400066 | 15954 | DvGlutMARCM-F347_seg1 | VGlut-Gal4 | F | 112377.53 | 1.00 |
| fru-M-400007 | 1778 | FruMARCM-M000295_seg002 | fru-Gal4 | M | 74992.92 | 0.55 |
| fru-F-500276 | 3773 | FruMARCM-F002397_seg002 | fru-Gal4 | F | 20081.57 | 0.19 |
| Cha-F-500178 | 3838 | ChaMARCM-F001513_seg001 | Cha-Gal4 | F | 41088.44 | 0.57 |
| Cha-F-000291 | 4176 | ChaMARCM-F001283_seg001 | Cha-Gal4 | F | 59700.11 | 0.55 |
| Cha-F-000361 | 4420 | ChaMARCM-F001467_seg003 | Cha-Gal4 | F | 38771.03 | 0.48 |
| Cha-F-000255 | 5173 | ChaMARCM-F001213_seg002 | Cha-Gal4 | F | 55702.61 | 0.51 |
| Cha-F-200135 | 5309 | ChaMARCM-F000691_seg001 | Cha-Gal4 | F | 44169.44 | 0.35 |
| Cha-F-400183 | 5469 | ChaMARCM-F000799_seg005 | Cha-Gal4 | F | 48578.64 | 0.61 |
| Cha-F-200189 | 5686 | ChaMARCM-F000945_seg001 | Cha-Gal4 | F | 63210.88 | 0.44 |
| VGlut-F-200587 | 5707 | DvGlutMARCM-F004417_seg003 | VGlut-Gal4 | F | 45083.63 | 0.39 |
| VGlut-F-400885 | 6458 | DvGlutMARCM-F004243_seg001 | VGlut-Gal4 | F | 77710.20 | 0.74 |
| Cha-F-600056 | 7583 | ChaMARCM-F000439_seg002 | Cha-Gal4 | F | 80204.32 | 0.62 |
| VGlut-F-000527 | 8957 | DvGlutMARCM-F003518_seg002 | VGlut-Gal4 | F | 80101.85 | 0.59 |
| Cha-F-700015 | 10167 | ChaMARCM-F000098_seg002 | Cha-Gal4 | F | 43003.85 | 0.46 |
| Gad1-F-000012 | 10666 | GadMARCM-F000174_seg003 | Gad1-Gal4 | F | 56784.33 | 0.65 |
| VGlut-F-000469 | 10842 | DvGlutMARCM-F002880_seg001 | VGlut-Gal4 | F | 73429.61 | 0.71 |
| Gad1-F-200066 | 11251 | GadMARCM-F000231_seg001 | Gad1-Gal4 | F | 64352.76 | 0.66 |
| VGlut-F-200397 | 11765 | DvGlutMARCM-F002336_seg001 | VGlut-Gal4 | F | 68673.86 | 0.63 |
| VGlut-F-400580 | 12286 | DvGlutMARCM-F002742_seg001 | VGlut-Gal4 | F | 78998.25 | 0.74 |