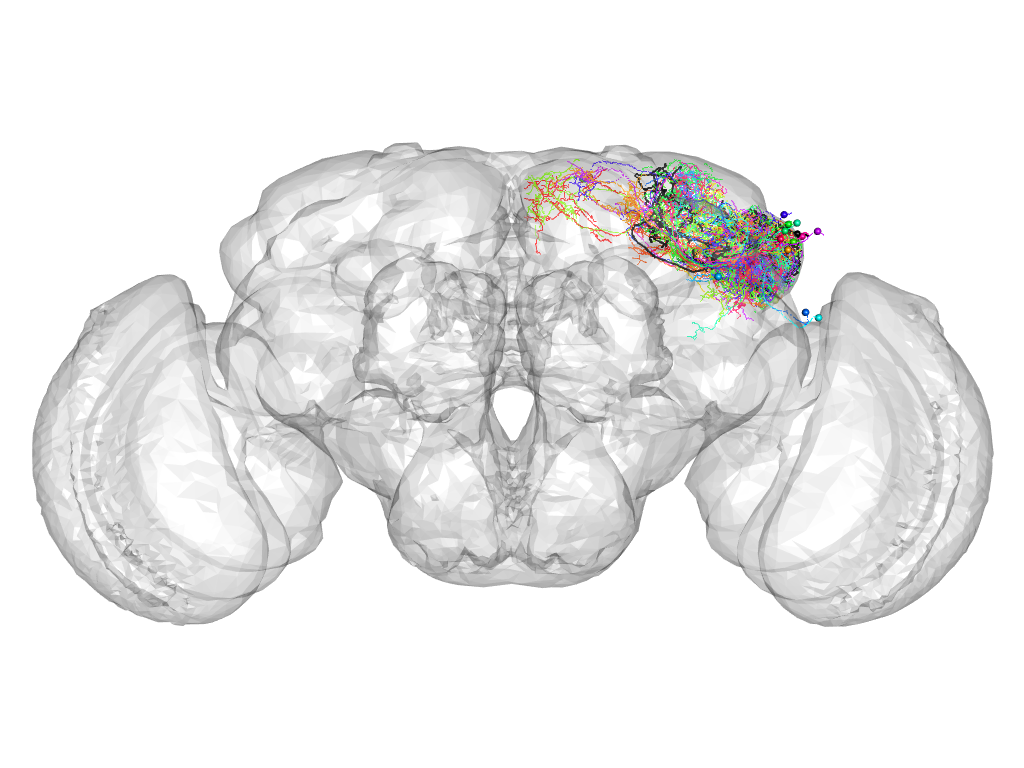
Cluster 339
Part of supercluster XVII
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (Cha-F-100223), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 136 | TH-M-000013 | XVII | 0.68 |
| 662 | Tdc2-F-400026 | XIII | 0.51 |
| 2 | fru-M-200253 | XVII | 0.41 |
| 360 | Cha-F-400179 | XVII | 0.39 |
| 366 | Cha-F-200185 | XVII | 0.38 |
Cluster composition
This cluster has 21 neurons. The exemplar of this cluster (Cha-F-100223) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| Cha-F-100223 | 5065 | ChaMARCM-F001138_seg001 | Cha-Gal4 | F | 64212.48 | 1.00 |
| fru-M-000077 | 496 | FruMARCM-M001592_seg001 | fru-Gal4 | M | 36843.97 | 0.48 |
| Cha-F-400240 | 3868 | ChaMARCM-F001535_seg001 | Cha-Gal4 | F | 27610.32 | 0.47 |
| Gad1-F-900037 | 3953 | GadMARCM-F000581_seg001 | Gad1-Gal4 | F | 26339.47 | 0.45 |
| Gad1-F-300135 | 3966 | GadMARCM-F000590_seg001 | Gad1-Gal4 | F | 31581.98 | 0.56 |
| Cha-F-100278 | 4239 | ChaMARCM-F001325_seg001 | Cha-Gal4 | F | 31842.80 | 0.62 |
| fru-F-000098 | 4640 | FruMARCM-F001807_seg001 | fru-Gal4 | F | 38058.17 | 0.49 |
| Cha-F-300272 | 5037 | ChaMARCM-F001121_seg002 | Cha-Gal4 | F | 36386.51 | 0.63 |
| VGlut-F-200319 | 6089 | DvGlutMARCM-F1730_seg1 | VGlut-Gal4 | F | 45068.39 | 0.69 |
| VGlut-F-700522 | 6447 | DvGlutMARCM-F004232_seg001 | VGlut-Gal4 | F | 33541.38 | 0.61 |
| VGlut-F-400863 | 6941 | DvGlutMARCM-F004095_seg001 | VGlut-Gal4 | F | 31286.92 | 0.56 |
| Cha-F-500105 | 7562 | ChaMARCM-F000411_seg003 | Cha-Gal4 | F | 29786.00 | 0.58 |
| Cha-F-700110 | 7584 | ChaMARCM-F000440_seg001 | Cha-Gal4 | F | 18719.13 | 0.48 |
| Cha-F-000048 | 8326 | ChaMARCM-F000284_seg003 | Cha-Gal4 | F | 23269.33 | 0.49 |
| Cha-F-300025 | 10178 | ChaMARCM-F000107_seg001 | Cha-Gal4 | F | 19231.01 | 0.37 |
| Gad1-F-500014 | 10498 | GadMARCM-F000152_seg001 | Gad1-Gal4 | F | 25404.67 | 0.44 |
| Gad1-F-500023 | 10507 | GadMARCM-F000155_seg002 | Gad1-Gal4 | F | 18582.78 | 0.43 |
| Gad1-F-200039 | 10677 | GadMARCM-F000182_seg001 | Gad1-Gal4 | F | 22785.37 | 0.37 |
| Gad1-F-100003 | 13168 | GadMARCM-F005_seg1 | Gad1-Gal4 | F | 26855.23 | 0.53 |
| VGlut-F-400267 | 14641 | DvGlutMARCM-F896_seg2 | VGlut-Gal4 | F | 33990.00 | 0.61 |
| VGlut-F-500032 | 15759 | DvGlutMARCM-F166_seg1 | VGlut-Gal4 | F | 31221.89 | 0.56 |