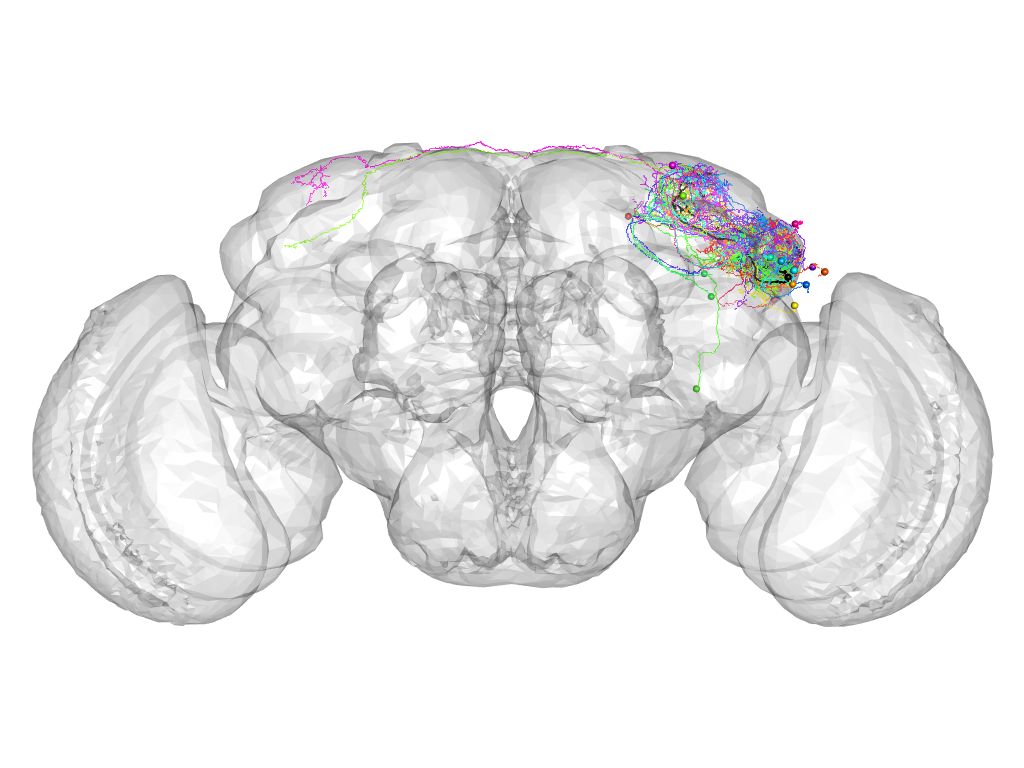
Cluster 366
Part of supercluster XVII
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (Cha-F-200185), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 339 | Cha-F-100223 | XVII | 0.69 |
| 136 | TH-M-000013 | XVII | 0.69 |
| 662 | Tdc2-F-400026 | XIII | 0.61 |
| 455 | VGlut-F-600764 | XVII | 0.60 |
| 360 | Cha-F-400179 | XVII | 0.58 |
Cluster composition
This cluster has 21 neurons. The exemplar of this cluster (Cha-F-200185) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| Cha-F-200185 | 5646 | ChaMARCM-F000922_seg002 | Cha-Gal4 | F | 34087.96 | 1.00 |
| Gad1-F-900054 | 4036 | GadMARCM-F000641_seg001 | Gad1-Gal4 | F | 22609.08 | 0.62 |
| Gad1-F-800082 | 4075 | GadMARCM-F000665_seg001 | Gad1-Gal4 | F | 22099.63 | 0.66 |
| Gad1-F-200144 | 4094 | GadMARCM-F000684_seg001 | Gad1-Gal4 | F | 24480.95 | 0.70 |
| Gad1-F-000096 | 4108 | GadMARCM-F000693_seg002 | Gad1-Gal4 | F | 22571.91 | 0.62 |
| Gad1-F-200150 | 4114 | GadMARCM-F000699_seg001 | Gad1-Gal4 | F | 17225.06 | 0.54 |
| Gad1-F-000108 | 4129 | GadMARCM-F000715_seg001 | Gad1-Gal4 | F | 19166.00 | 0.41 |
| Cha-F-000278 | 4151 | ChaMARCM-F001266_seg001 | Cha-Gal4 | F | 24336.31 | 0.66 |
| Cha-F-000317 | 4267 | ChaMARCM-F001346_seg002 | Cha-Gal4 | F | 22153.62 | 0.60 |
| Cha-F-100141 | 4886 | ChaMARCM-F001003_seg001 | Cha-Gal4 | F | 20260.80 | 0.56 |
| Cha-F-000258 | 5188 | ChaMARCM-F001225_seg001 | Cha-Gal4 | F | 21795.54 | 0.60 |
| Cha-F-400151 | 5289 | ChaMARCM-F000679_seg001 | Cha-Gal4 | F | 22991.11 | 0.70 |
| Cha-F-000158 | 5586 | ChaMARCM-F000878_seg002 | Cha-Gal4 | F | 19098.11 | 0.55 |
| Cha-F-300258 | 5672 | ChaMARCM-F000937_seg001 | Cha-Gal4 | F | 20897.02 | 0.57 |
| Cha-F-100068 | 7111 | ChaMARCM-F000585_seg001 | Cha-Gal4 | F | 20292.94 | 0.52 |
| Cha-F-600051 | 7539 | ChaMARCM-F000393_seg003 | Cha-Gal4 | F | 18745.40 | 0.50 |
| Cha-F-700026 | 8080 | ChaMARCM-F000145_seg001 | Cha-Gal4 | F | 21900.49 | 0.60 |
| Gad1-F-100025 | 11271 | GadMARCM-F000247_seg002 | Gad1-Gal4 | F | 21407.49 | 0.56 |
| Cha-F-100001 | 11567 | ChaMARCM-F000002_seg001 | Cha-Gal4 | F | 12312.30 | 0.21 |
| VGlut-F-600424 | 12295 | DvGlutMARCM-F002751_seg001 | VGlut-Gal4 | F | 23086.00 | 0.62 |
| VGlut-F-400394 | 15246 | DvGlutMARCM-F1567_seg1 | VGlut-Gal4 | F | 19628.16 | 0.54 |