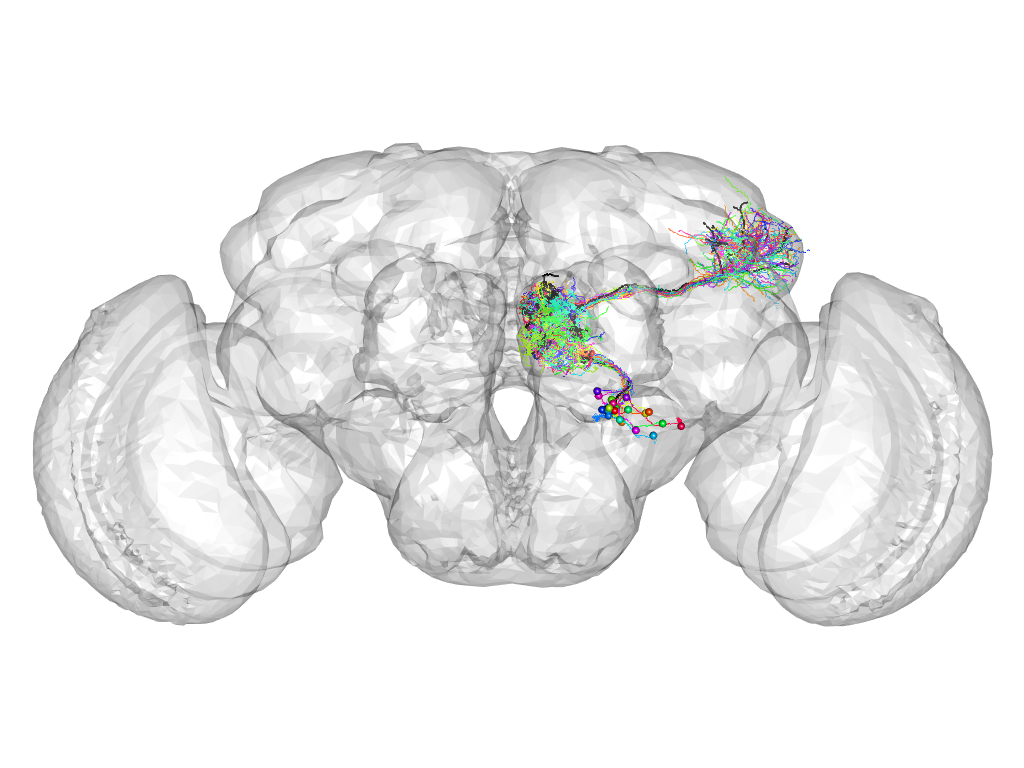
Cluster 911
Part of supercluster XI
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (VGlut-F-400172), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 662 | Tdc2-F-400026 | XIII | 0.66 |
| 375 | VGlut-F-500848 | XI | 0.55 |
| 806 | Trh-F-100085 | XII | 0.54 |
| 144 | Trh-M-400113 | XII | 0.47 |
| 201 | Trh-M-100007 | XII | 0.46 |
Cluster composition
This cluster has 23 neurons. The exemplar of this cluster (VGlut-F-400172) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| VGlut-F-400172 | 14434 | DvGlutMARCM-F725_seg1 | VGlut-Gal4 | F | 45169.69 | 1.00 |
| fru-M-800089 | 61 | FruMARCM-M002255_seg002 | fru-Gal4 | M | 30655.74 | 0.69 |
| Trh-M-500165 | 1997 | TPHMARCM-M001532_seg001 | Trh-Gal4 | M | 29765.13 | 0.69 |
| Trh-M-600114 | 2220 | TPHMARCM-M001710_seg002 | Trh-Gal4 | M | 29868.49 | 0.68 |
| Trh-M-600013 | 3594 | TPHMARCM-537M_seg2 | Trh-Gal4 | M | 29412.69 | 0.70 |
| VGlut-F-700559 | 5731 | DvGlutMARCM-F004439_seg001 | VGlut-Gal4 | F | 30419.70 | 0.67 |
| VGlut-F-300636 | 6559 | DvGlutMARCM-F004321_seg001 | VGlut-Gal4 | F | 16887.09 | 0.49 |
| VGlut-F-500817 | 6600 | DvGlutMARCM-F004358_seg001 | VGlut-Gal4 | F | 29113.85 | 0.67 |
| VGlut-F-700473 | 6829 | DvGlutMARCM-F004012_seg001 | VGlut-Gal4 | F | 27066.42 | 0.61 |
| VGlut-F-700420 | 8003 | DvGlutMARCM-F003823_seg001 | VGlut-Gal4 | F | 30344.94 | 0.69 |
| VGlut-F-700428 | 8028 | DvGlutMARCM-F003841_seg001 | VGlut-Gal4 | F | 28626.57 | 0.68 |
| VGlut-F-600669 | 8738 | DvGlutMARCM-F003565_seg002 | VGlut-Gal4 | F | 30745.22 | 0.68 |
| VGlut-F-500687 | 8950 | DvGlutMARCM-F003514_seg001 | VGlut-Gal4 | F | 27192.16 | 0.66 |
| VGlut-F-700218 | 10788 | DvGlutMARCM-F002842_seg002 | VGlut-Gal4 | F | 27327.34 | 0.61 |
| Trh-F-600095 | 12448 | TPHMARCM-1242F_seg2 | Trh-Gal4 | F | 24946.26 | 0.62 |
| Trh-F-600066 | 12728 | TPHMARCM-930F_seg1 | Trh-Gal4 | F | 28647.70 | 0.66 |
| Trh-F-800002 | 13267 | TPHMARCM-754F_seg1 | Trh-Gal4 | F | 30446.80 | 0.68 |
| Trh-F-500049 | 14154 | TPHMARCM-578F_seg1 | Trh-Gal4 | F | 29430.99 | 0.68 |
| Trh-F-500059 | 14195 | TPHMARCM-637F_seg1 | Trh-Gal4 | F | 30196.93 | 0.70 |
| VGlut-F-400167 | 14429 | DvGlutMARCM-F722_seg1 | VGlut-Gal4 | F | 28742.51 | 0.69 |
| VGlut-F-400263 | 14633 | DvGlutMARCM-F889_seg1 | VGlut-Gal4 | F | 27866.71 | 0.65 |
| VGlut-F-500205 | 15275 | DvGlutMARCM-F1602_seg1 | VGlut-Gal4 | F | 30206.45 | 0.69 |
| VGlut-F-700022 | 15311 | DvGlutMARCM-F1647_seg1 | VGlut-Gal4 | F | 29818.63 | 0.65 |