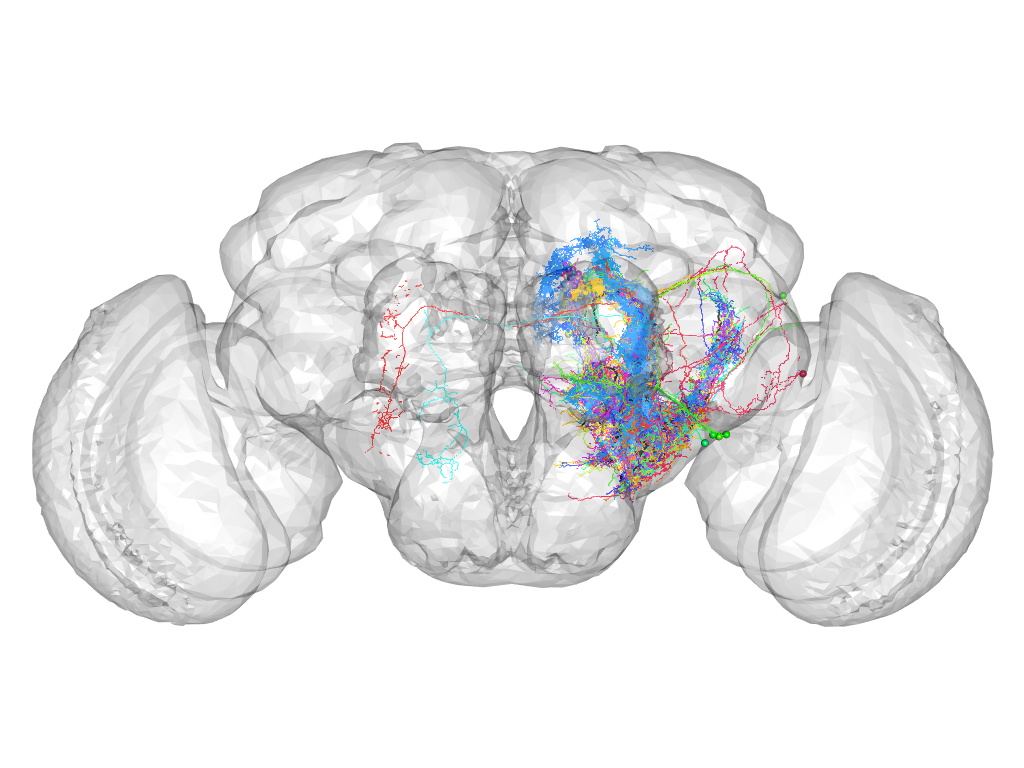
Cluster 855
Part of supercluster VIII
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (TH-F-200057), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 187 | TH-M-400003 | VIII | 0.72 |
| 921 | VGlut-F-300181 | VIII | 0.60 |
| 667 | TH-F-300071 | XIV | 0.58 |
| 988 | VGlut-F-200005 | X | 0.54 |
| 406 | fru-F-000053 | VII | 0.54 |
| 753 | VGlut-F-600178 | XIV | 0.53 |
| 191 | TH-M-200018 | XIV | 0.50 |
Cluster composition
This cluster has 56 neurons. The exemplar of this cluster (TH-F-200057) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| TH-F-200057 | 13506 | THMARCM-383F_seg1 | TH-Gal4 | F | 128288.28 | 1.00 |
| fru-M-100201 | 904 | FruMARCM-M002169_seg002 | fru-Gal4 | M | 83325.92 | 0.54 |
| TH-M-200013 | 944 | THMARCM-259M_seg2 | TH-Gal4 | M | 97043.62 | 0.77 |
| TH-M-200031 | 1331 | THMARCM-318M_seg1 | TH-Gal4 | M | 89374.88 | 0.73 |
| TH-M-300016 | 3107 | THMARCM-092M_seg1 | TH-Gal4 | M | 84484.06 | 0.65 |
| TH-M-200001 | 3135 | THMARCM-176M_seg1 | TH-Gal4 | M | 90490.58 | 0.73 |
| TH-M-100026 | 3157 | THMARCM-207M_seg1 | TH-Gal4 | M | 92209.31 | 0.73 |
| TH-M-200015 | 3172 | THMARCM-261M_seg1 | TH-Gal4 | M | 91513.51 | 0.73 |
| TH-M-100040 | 3185 | THMARCM-280M_seg2 | TH-Gal4 | M | 97412.66 | 0.72 |
| TH-M-100047 | 3201 | THMARCM-319M_seg1 | TH-Gal4 | M | 93643.17 | 0.74 |
| TH-M-100048 | 3202 | THMARCM-319M_seg2 | TH-Gal4 | M | 89445.92 | 0.73 |
| TH-M-100052 | 3213 | THMARCM-342M_seg1 | TH-Gal4 | M | 89926.68 | 0.71 |
| TH-M-200043 | 3224 | THMARCM-358M_seg2 | TH-Gal4 | M | 90315.09 | 0.68 |
| TH-M-200059 | 3240 | THMARCM-395M_seg2 | TH-Gal4 | M | 91202.16 | 0.69 |
| TH-M-200063 | 3245 | THMARCM-401M_seg2 | TH-Gal4 | M | 91803.30 | 0.71 |
| TH-M-200070 | 3257 | THMARCM-436M_seg1 | TH-Gal4 | M | 90674.95 | 0.71 |
| TH-M-000062 | 3368 | THMARCM-781M_seg1 | TH-Gal4 | M | 87666.18 | 0.73 |
| Cha-F-200275 | 4141 | ChaMARCM-F001258_seg001 | Cha-Gal4 | F | 70822.03 | 0.52 |
| Cha-F-200310 | 4253 | ChaMARCM-F001336_seg001 | Cha-Gal4 | F | 44927.46 | 0.35 |
| fru-F-600091 | 4503 | FruMARCM-F001605_seg001 | fru-Gal4 | F | 55631.38 | 0.56 |
| fru-F-500227 | 4702 | FruMARCM-F001964_seg001 | fru-Gal4 | F | 51489.91 | 0.51 |
| VGlut-F-200528 | 8716 | DvGlutMARCM-F003749_seg002 | VGlut-Gal4 | F | 87388.98 | 0.72 |
| VGlut-F-100301 | 8903 | DvGlutMARCM-F003478_seg002 | VGlut-Gal4 | F | 88257.79 | 0.70 |
| fru-F-500027 | 9908 | FruMARCM-F000494_seg001 | fru-Gal4 | F | 44264.77 | 0.51 |
| TH-F-100015 | 10467 | THMARCM-209F_seg2 | TH-Gal4 | F | 89869.74 | 0.71 |
| TH-F-200042 | 10474 | THMARCM-345F_seg1 | TH-Gal4 | F | 93124.23 | 0.73 |
| TH-F-100112 | 10493 | THMARCM-800F_seg2 | TH-Gal4 | F | 88616.89 | 0.73 |
| TH-F-000093 | 10494 | THMARCM-811F_seg1 | TH-Gal4 | F | 88221.94 | 0.72 |
| fru-F-200019 | 11515 | FruMARCM-F000176_seg001 | fru-Gal4 | F | 79216.68 | 0.49 |
| TH-F-300010 | 13387 | THMARCM-155F_seg3 | TH-Gal4 | F | 91122.72 | 0.74 |
| TH-F-300016 | 13392 | THMARCM-160F_seg2 | TH-Gal4 | F | 89695.38 | 0.71 |
| TH-F-100013 | 13407 | THMARCM-206F_seg1 | TH-Gal4 | F | 86640.01 | 0.70 |
| TH-F-300034 | 13435 | THMARCM-250F_seg2 | TH-Gal4 | F | 103952.51 | 0.74 |
| TH-F-200015 | 13439 | THMARCM-265F_seg2 | TH-Gal4 | F | 103033.42 | 0.72 |
| TH-F-200016 | 13440 | THMARCM-266F_seg1 | TH-Gal4 | F | 102062.49 | 0.76 |
| TH-F-300036 | 13443 | THMARCM-276F_seg1 | TH-Gal4 | F | 98648.21 | 0.76 |
| TH-F-200028 | 13461 | THMARCM-298F_seg2 | TH-Gal4 | F | 90773.81 | 0.72 |
| TH-F-300039 | 13464 | THMARCM-300F_seg2 | TH-Gal4 | F | 92218.96 | 0.70 |
| TH-F-300041 | 13478 | THMARCM-325F_seg1 | TH-Gal4 | F | 90322.55 | 0.73 |
| TH-F-100040 | 13485 | THMARCM-337F_seg2 | TH-Gal4 | F | 97727.42 | 0.76 |
| TH-F-200044 | 13488 | THMARCM-349F_seg1 | TH-Gal4 | F | 87901.42 | 0.72 |
| TH-F-200060 | 13509 | THMARCM-384F_seg2 | TH-Gal4 | F | 91483.00 | 0.71 |
| TH-F-100042 | 13526 | THMARCM-409F_seg1 | TH-Gal4 | F | 92133.59 | 0.72 |
| TH-F-100043 | 13527 | THMARCM-409F_seg2 | TH-Gal4 | F | 90031.73 | 0.72 |
| TH-F-300064 | 13560 | THMARCM-461F_seg2 | TH-Gal4 | F | 92977.20 | 0.74 |
| TH-F-000024 | 13607 | THMARCM-550F_seg1 | TH-Gal4 | F | 87098.16 | 0.70 |
| TH-F-000034 | 13632 | THMARCM-619F-X2_seg2 | TH-Gal4 | F | 91191.33 | 0.71 |
| TH-F-100076 | 13643 | THMARCM-647F_seg | TH-Gal4 | F | 91709.98 | 0.72 |
| TH-F-000060 | 13682 | THMARCM-705F_seg1 | TH-Gal4 | F | 89806.80 | 0.71 |
| TH-F-200118 | 13701 | THMARCM-722F_seg2 | TH-Gal4 | F | 93687.07 | 0.74 |
| TH-F-100087 | 13712 | THMARCM-739F_seg2 | TH-Gal4 | F | 89811.09 | 0.68 |
| TH-F-000070 | 13714 | THMARCM-746F_seg1 | TH-Gal4 | F | 89836.71 | 0.73 |
| TH-F-000075 | 13724 | THMARCM-754F_seg1 | TH-Gal4 | F | 94012.92 | 0.75 |
| TH-F-000092 | 13759 | THMARCM-797F_seg1 | TH-Gal4 | F | 88468.16 | 0.70 |
| VGlut-F-000236 | 14711 | DvGlutMARCM-F970_seg1 | VGlut-Gal4 | F | 78129.84 | 0.51 |
| VGlut-F-100178 | 15355 | DvGlutMARCM-F1704_seg1 | VGlut-Gal4 | F | 85688.72 | 0.71 |