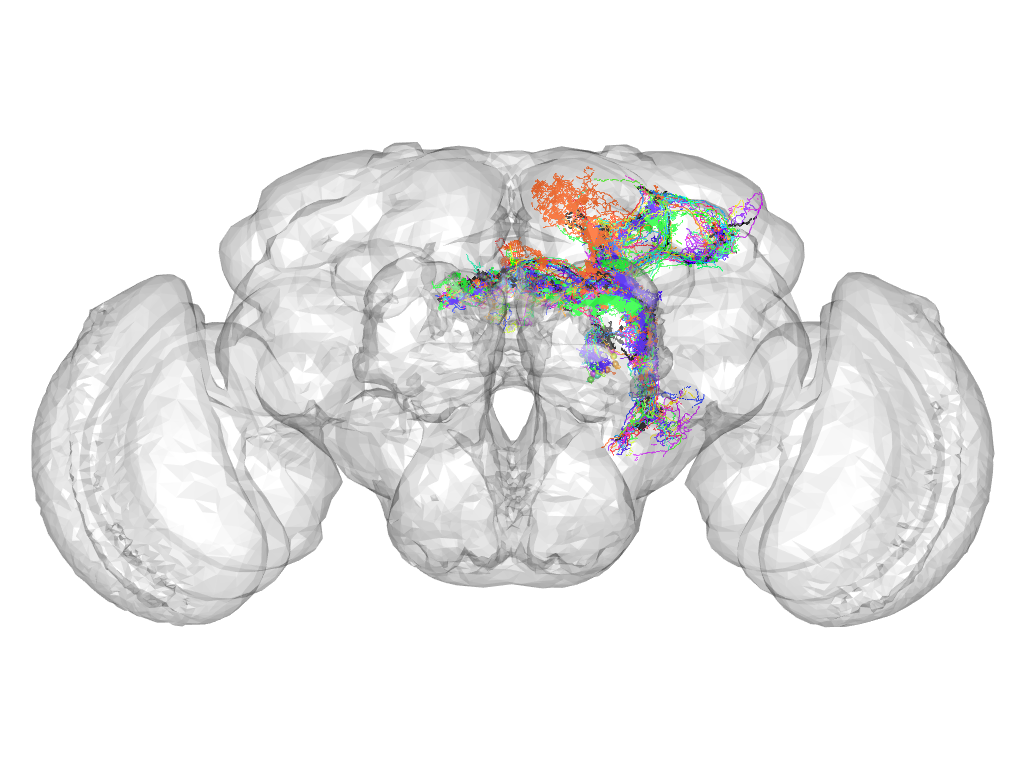
Cluster 191
Part of supercluster XIV
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (TH-M-200018), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 192 | TH-M-300027 | XIV | 0.61 |
| 1028 | VGlut-F-000067 | XIV | 0.55 |
| 667 | TH-F-300071 | XIV | 0.52 |
| 851 | TH-F-100029 | XIV | 0.43 |
| 204 | Trh-M-100025 | XIV | 0.40 |
Cluster composition
This cluster has 21 neurons. The exemplar of this cluster (TH-M-200018) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| TH-M-200018 | 3175 | THMARCM-263M_seg1 | TH-Gal4 | M | 140212.81 | 1.00 |
| fru-M-200269 | 90 | FruMARCM-M002289_seg001 | fru-Gal4 | M | 98594.13 | 0.71 |
| TH-M-200012 | 943 | THMARCM-259M_seg1 | TH-Gal4 | M | 107181.34 | 0.70 |
| fru-M-500131 | 1076 | FruMARCM-M001252_seg001 | fru-Gal4 | M | 95487.92 | 0.69 |
| fru-M-200102 | 1481 | FruMARCM-M001038_seg001 | fru-Gal4 | M | 96116.93 | 0.71 |
| fru-M-100016 | 2471 | FruMARCM-M000227_seg001 | fru-Gal4 | M | 97191.94 | 0.71 |
| TH-M-200020 | 2587 | THMARCM-277M_seg1 | TH-Gal4 | M | 94077.16 | 0.72 |
| TH-M-300003 | 3085 | THMARCM-062M_seg1 | TH-Gal4 | M | 97989.54 | 0.70 |
| TH-M-200019 | 3178 | THMARCM-270M_seg1 | TH-Gal4 | M | 101848.55 | 0.71 |
| TH-M-200040 | 3215 | THMARCM-344M_seg1 | TH-Gal4 | M | 95371.38 | 0.71 |
| TH-M-200067 | 3253 | THMARCM-421M_seg1 | TH-Gal4 | M | 96090.35 | 0.72 |
| TH-M-200068 | 3254 | THMARCM-426M_seg1 | TH-Gal4 | M | 96221.36 | 0.73 |
| TH-M-200093 | 3320 | THMARCM-568M_seg1 | TH-Gal4 | M | 95371.89 | 0.71 |
| TH-M-300062 | 3362 | THMARCM-772M_seg1 | TH-Gal4 | M | 97289.01 | 0.74 |
| fru-F-200031 | 9827 | FruMARCM-F000394_seg001 | fru-Gal4 | F | 92833.81 | 0.68 |
| TH-F-200014 | 13438 | THMARCM-265F_seg1 | TH-Gal4 | F | 109984.04 | 0.75 |
| TH-F-300043 | 13473 | THMARCM-313F_seg1 | TH-Gal4 | F | 95087.74 | 0.70 |
| TH-F-200047 | 13491 | THMARCM-351F_seg1 | TH-Gal4 | F | 97466.48 | 0.69 |
| TH-F-200055 | 13504 | THMARCM-381F_seg1 | TH-Gal4 | F | 96462.81 | 0.71 |
| TH-F-200106 | 13668 | THMARCM-682F_seg | TH-Gal4 | F | 96447.72 | 0.73 |
| TH-F-200126 | 13773 | THMARCM-825F_seg1 | TH-Gal4 | F | 96256.30 | 0.73 |