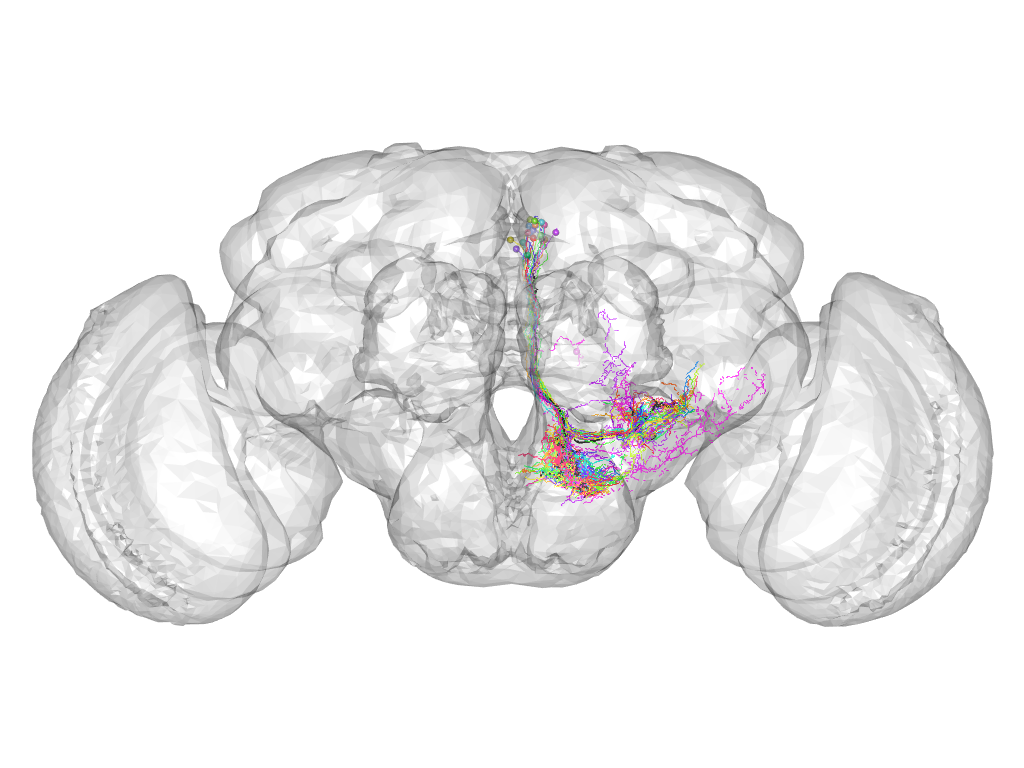
Cluster 765
Part of supercluster VIII
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (VGlut-F-500530), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 441 | VGlut-F-600718 | VIII | 0.73 |
| 690 | VGlut-F-400061 | VIII | 0.72 |
| 652 | Tdc2-F-200010 | VIII | 0.60 |
| 430 | VGlut-F-500830 | VIII | 0.51 |
| 751 | VGlut-F-400648 | VIII | 0.50 |
Cluster composition
This cluster has 20 neurons. The exemplar of this cluster (VGlut-F-500530) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| VGlut-F-500530 | 11905 | DvGlutMARCM-F002441_seg003 | VGlut-Gal4 | F | 17072.46 | 1.00 |
| fru-M-100159 | 529 | FruMARCM-M001649_seg002 | fru-Gal4 | M | 11850.66 | 0.66 |
| fru-M-000056 | 1141 | FruMARCM-M001334_seg001 | fru-Gal4 | M | 11236.29 | 0.64 |
| fru-M-500139 | 1287 | FruMARCM-M001475_seg002 | fru-Gal4 | M | 12585.15 | 0.69 |
| fru-M-400086 | 1461 | FruMARCM-M001023_seg002 | fru-Gal4 | M | 11346.47 | 0.64 |
| fru-M-500035 | 1958 | FruMARCM-M000600_seg001 | fru-Gal4 | M | 12108.28 | 0.58 |
| fru-F-000040 | 6170 | FruMARCM-F001107_seg001 | fru-Gal4 | F | 11443.42 | 0.65 |
| VGlut-F-700505 | 7046 | DvGlutMARCM-F004176_seg002 | VGlut-Gal4 | F | 12155.04 | 0.70 |
| VGlut-F-500611 | 9188 | DvGlutMARCM-F003256_seg002 | VGlut-Gal4 | F | 12498.47 | 0.64 |
| VGlut-F-500597 | 9378 | DvGlutMARCM-F003209_seg001 | VGlut-Gal4 | F | 12413.00 | 0.73 |
| VGlut-F-500027 | 10237 | DvGlutMARCM-F163_seg1 | VGlut-Gal4 | F | 12670.61 | 0.71 |
| VGlut-F-700077 | 12084 | DvGlutMARCM-F002591_seg003 | VGlut-Gal4 | F | 12701.09 | 0.74 |
| VGlut-F-200349 | 12956 | DvGlutMARCM-F1987_seg2 | VGlut-Gal4 | F | 11728.87 | 0.63 |
| VGlut-F-400500 | 13098 | DvGlutMARCM-F2121_seg1 | VGlut-Gal4 | F | 12681.41 | 0.74 |
| VGlut-F-400256 | 14626 | DvGlutMARCM-F883_seg2 | VGlut-Gal4 | F | 12901.91 | 0.75 |
| VGlut-F-300187 | 14710 | DvGlutMARCM-F968_seg1 | VGlut-Gal4 | F | 12399.78 | 0.69 |
| VGlut-F-400326 | 14808 | DvGlutMARCM-F1155_seg1 | VGlut-Gal4 | F | 11039.63 | 0.47 |
| VGlut-F-000369 | 15190 | DvGlutMARCM-F1518_seg2 | VGlut-Gal4 | F | 9371.56 | 0.31 |
| VGlut-F-500194 | 15229 | DvGlutMARCM-F1551_seg1 | VGlut-Gal4 | F | 12825.77 | 0.72 |
| VGlut-F-500023 | 15735 | DvGlutMARCM-F145_seg1 | VGlut-Gal4 | F | 10700.87 | 0.61 |