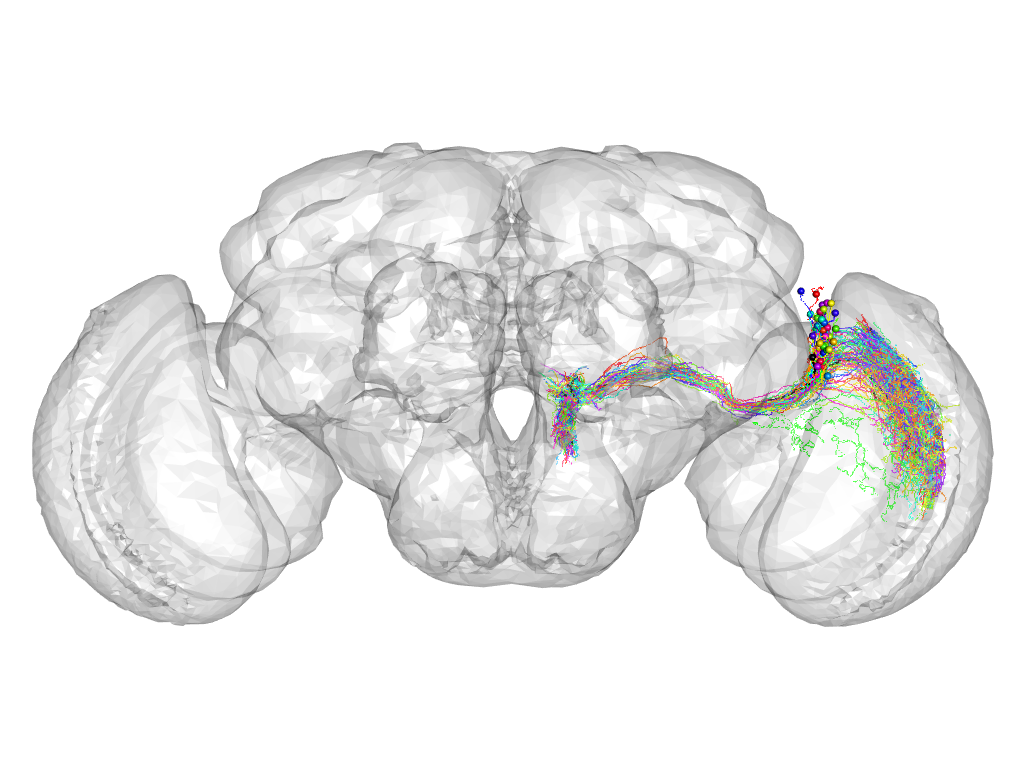
Cluster 57
Part of supercluster III
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (fru-M-600086), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 661 | Tdc2-F-000027 | VII | 0.56 |
| 759 | VGlut-F-600257 | III | 0.56 |
| 660 | Tdc2-F-200049 | I | 0.50 |
| 468 | VGlut-F-600670 | III | 0.49 |
| 175 | Trh-M-700055 | III | 0.45 |
Cluster composition
This cluster has 50 neurons. The exemplar of this cluster (fru-M-600086) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| fru-M-600086 | 697 | FruMARCM-M001900_seg001 | fru-Gal4 | M | 23085.97 | 1.00 |
| fru-M-500227 | 39 | FruMARCM-M002235_seg001 | fru-Gal4 | M | 15195.10 | 0.65 |
| fru-M-300257 | 123 | FruMARCM-M002318_seg001 | fru-Gal4 | M | 16863.00 | 0.72 |
| fru-M-900063 | 172 | FruMARCM-M002390_seg001 | fru-Gal4 | M | 15379.25 | 0.66 |
| fru-M-100211 | 288 | FruMARCM-M002513_seg001 | fru-Gal4 | M | 16644.16 | 0.72 |
| fru-M-100218 | 301 | FruMARCM-M002528_seg001 | fru-Gal4 | M | 16061.78 | 0.69 |
| fru-M-100224 | 328 | FruMARCM-M002550_seg001 | fru-Gal4 | M | 16283.29 | 0.70 |
| fru-M-100226 | 355 | FruMARCM-M002589_seg001 | fru-Gal4 | M | 15641.02 | 0.66 |
| fru-M-700104 | 414 | FruMARCM-M001839_seg001 | fru-Gal4 | M | 16769.84 | 0.71 |
| fru-M-900045 | 694 | FruMARCM-M001897_seg001 | fru-Gal4 | M | 14799.80 | 0.65 |
| fru-M-300136 | 1068 | FruMARCM-M001246_seg001 | fru-Gal4 | M | 16091.02 | 0.69 |
| fru-M-300142 | 1142 | FruMARCM-M001338_seg001 | fru-Gal4 | M | 15647.75 | 0.67 |
| fru-M-200178 | 1200 | FruMARCM-M001379_seg001 | fru-Gal4 | M | 14402.63 | 0.61 |
| fru-M-200183 | 1205 | FruMARCM-M001382_seg001 | fru-Gal4 | M | 15332.21 | 0.68 |
| fru-M-500077 | 1391 | FruMARCM-M000894_seg002 | fru-Gal4 | M | 14679.31 | 0.62 |
| fru-M-400068 | 1419 | FruMARCM-M000933_seg001 | fru-Gal4 | M | 16259.84 | 0.67 |
| fru-M-200096 | 1447 | FruMARCM-M001016_seg001 | fru-Gal4 | M | 14511.87 | 0.66 |
| fru-M-100087 | 1525 | FruMARCM-M001067_seg001 | fru-Gal4 | M | 15038.11 | 0.46 |
| fru-M-100089 | 1527 | FruMARCM-M001068_seg001 | fru-Gal4 | M | 16753.19 | 0.71 |
| fru-M-000019 | 1563 | FruMARCM-M000632_seg002 | fru-Gal4 | M | 16100.91 | 0.69 |
| fru-M-300080 | 1634 | FruMARCM-M000773_seg001 | fru-Gal4 | M | 16486.21 | 0.71 |
| fru-M-100063 | 1644 | FruMARCM-M000784_seg001 | fru-Gal4 | M | 14789.93 | 0.67 |
| fru-M-300093 | 1714 | FruMARCM-M000866_seg001 | fru-Gal4 | M | 15255.93 | 0.68 |
| fru-M-800018 | 1846 | FruMARCM-M000357_seg001 | fru-Gal4 | M | 15519.15 | 0.68 |
| fru-M-700025 | 1864 | FruMARCM-M000410_seg001 | fru-Gal4 | M | 15009.96 | 0.60 |
| fru-M-100010 | 2380 | FruMARCM-M000109_seg001 | fru-Gal4 | M | 15201.34 | 0.67 |
| fru-M-800006 | 2390 | FruMARCM-M000117_seg001 | fru-Gal4 | M | 15454.62 | 0.69 |
| fru-M-700015 | 2435 | FruMARCM-M000152_seg001 | fru-Gal4 | M | 16418.67 | 0.70 |
| fru-M-400004 | 2457 | FruMARCM-M000216_seg001 | fru-Gal4 | M | 15911.89 | 0.69 |
| fru-M-300031 | 2463 | FruMARCM-M000221_seg001 | fru-Gal4 | M | 14862.69 | 0.67 |
| fru-M-100019 | 2474 | FruMARCM-M000230_seg001 | fru-Gal4 | M | 16056.01 | 0.70 |
| fru-M-800015 | 2485 | FruMARCM-M000239_seg001 | fru-Gal4 | M | 15608.21 | 0.69 |
| fru-F-300120 | 3709 | FruMARCM-F002201_seg001 | fru-Gal4 | F | 15549.64 | 0.72 |
| fru-F-200153 | 3718 | FruMARCM-F002214_seg001 | fru-Gal4 | F | 15030.87 | 0.66 |
| fru-F-300092 | 4511 | FruMARCM-F001621_seg001 | fru-Gal4 | F | 15003.38 | 0.67 |
| fru-F-700157 | 4548 | FruMARCM-F001658_seg001 | fru-Gal4 | F | 15762.68 | 0.67 |
| fru-F-800052 | 4626 | FruMARCM-F001756_seg001 | fru-Gal4 | F | 15994.90 | 0.70 |
| fru-F-300111 | 4746 | FruMARCM-F002068_seg001 | fru-Gal4 | F | 15467.68 | 0.68 |
| fru-F-000111 | 4774 | FruMARCM-F002092_seg001 | fru-Gal4 | F | 16456.63 | 0.71 |
| fru-F-900027 | 4792 | FruMARCM-F002115_seg001 | fru-Gal4 | F | 15731.02 | 0.68 |
| fru-F-200085 | 6253 | FruMARCM-F001271_seg001 | fru-Gal4 | F | 15666.75 | 0.69 |
| fru-F-100050 | 6279 | FruMARCM-F001309_seg001 | fru-Gal4 | F | 14809.35 | 0.66 |
| fru-F-100051 | 6288 | FruMARCM-F001384_seg001 | fru-Gal4 | F | 14666.60 | 0.67 |
| fru-F-200101 | 6347 | FruMARCM-F001498_seg001 | fru-Gal4 | F | 16021.55 | 0.67 |
| fru-F-300054 | 7373 | FruMARCM-F001001_seg001 | fru-Gal4 | F | 15321.51 | 0.67 |
| fru-F-700059 | 7786 | FruMARCM-F000681_seg001 | fru-Gal4 | F | 16367.83 | 0.71 |
| fru-F-000012 | 9737 | FruMARCM-F000247_seg001 | fru-Gal4 | F | 14539.94 | 0.68 |
| fru-F-400057 | 9804 | FruMARCM-F000362_seg004 | fru-Gal4 | F | 15162.73 | 0.65 |
| fru-F-600005 | 11512 | FruMARCM-F000173_seg001 | fru-Gal4 | F | 15385.81 | 0.70 |
| fru-F-700013 | 11520 | FruMARCM-F000179_seg001 | fru-Gal4 | F | 13848.37 | 0.65 |