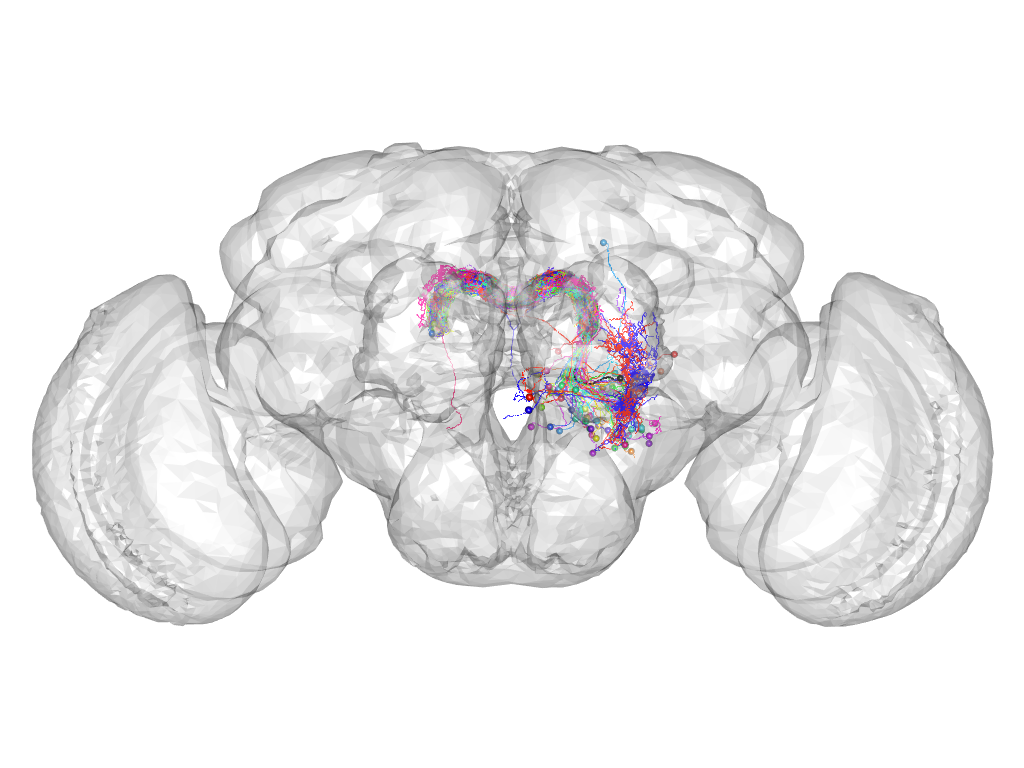
Cluster 527
Part of supercluster XIV
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (Cha-F-700075), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 1004 | VGlut-F-100016 | VI | 0.21 |
| 348 | Cha-F-200254 | VI | 0.20 |
| 1011 | VGlut-F-600010 | VI | 0.19 |
| 110 | fru-M-400100 | VII | 0.12 |
| 406 | fru-F-000053 | VII | 0.11 |
Cluster composition
This cluster has 79 neurons. The exemplar of this cluster (Cha-F-700075) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| Cha-F-700075 | 8235 | ChaMARCM-F000232_seg001 | Cha-Gal4 | F | 13621.52 | 1.00 |
| Trh-M-300080 | 2965 | TPHMARCM-906M_seg1 | Trh-Gal4 | M | -1252.04 | 0.27 |
| TH-M-000034 | 3057 | THMARCM-605M_seg | TH-Gal4 | M | 10475.13 | 0.50 |
| Gad1-F-400111 | 3938 | GadMARCM-F000568_seg001 | Gad1-Gal4 | F | 10937.88 | 0.80 |
| Cha-F-300305 | 4434 | ChaMARCM-F001478_seg002 | Cha-Gal4 | F | 10732.63 | 0.77 |
| Cha-F-200125 | 5267 | ChaMARCM-F000665_seg001 | Cha-Gal4 | F | 6582.34 | 0.55 |
| Cha-F-200130 | 5296 | ChaMARCM-F000683_seg001 | Cha-Gal4 | F | 10214.78 | 0.75 |
| Cha-F-300196 | 5301 | ChaMARCM-F000685_seg002 | Cha-Gal4 | F | 10426.13 | 0.74 |
| Cha-F-200148 | 5383 | ChaMARCM-F000747_seg001 | Cha-Gal4 | F | 10595.40 | 0.78 |
| Cha-F-300236 | 5516 | ChaMARCM-F000830_seg001 | Cha-Gal4 | F | 9944.62 | 0.72 |
| Cha-F-200165 | 5533 | ChaMARCM-F000840_seg002 | Cha-Gal4 | F | 10452.78 | 0.78 |
| VGlut-F-700566 | 5751 | DvGlutMARCM-F004456_seg001 | VGlut-Gal4 | F | 10296.97 | 0.78 |
| VGlut-F-500370 | 6100 | DvGlutMARCM-F2164_seg2 | VGlut-Gal4 | F | 10127.53 | 0.76 |
| VGlut-F-400888 | 6485 | DvGlutMARCM-F004263_seg001 | VGlut-Gal4 | F | 9790.29 | 0.74 |
| VGlut-F-000653 | 6496 | DvGlutMARCM-F004275_seg001 | VGlut-Gal4 | F | 10443.81 | 0.76 |
| VGlut-F-500797 | 6502 | DvGlutMARCM-F004281_seg001 | VGlut-Gal4 | F | 10089.86 | 0.75 |
| VGlut-F-600784 | 6555 | DvGlutMARCM-F004317_seg001 | VGlut-Gal4 | F | 9964.15 | 0.72 |
| VGlut-F-200570 | 6596 | DvGlutMARCM-F004355_seg001 | VGlut-Gal4 | F | 10108.39 | 0.75 |
| VGlut-F-800282 | 6602 | DvGlutMARCM-F004360_seg001 | VGlut-Gal4 | F | 10293.39 | 0.76 |
| VGlut-F-000619 | 6834 | DvGlutMARCM-F004017_seg001 | VGlut-Gal4 | F | 10251.97 | 0.77 |
| VGlut-F-600744 | 6903 | DvGlutMARCM-F004059_seg002 | VGlut-Gal4 | F | 8673.42 | 0.70 |
| VGlut-F-000634 | 6948 | DvGlutMARCM-F004102_seg001 | VGlut-Gal4 | F | 9590.73 | 0.73 |
| VGlut-F-800255 | 6969 | DvGlutMARCM-F004116_seg001 | VGlut-Gal4 | F | 9989.63 | 0.76 |
| VGlut-F-600765 | 6982 | DvGlutMARCM-F004128_seg001 | VGlut-Gal4 | F | 9290.38 | 0.73 |
| VGlut-F-500773 | 6997 | DvGlutMARCM-F004139_seg001 | VGlut-Gal4 | F | 10079.54 | 0.75 |
| Cha-F-900016 | 7469 | ChaMARCM-F000346_seg001 | Cha-Gal4 | F | 10029.29 | 0.73 |
| Cha-F-600063 | 7619 | ChaMARCM-F000461_seg001 | Cha-Gal4 | F | 10415.12 | 0.76 |
| Cha-F-600066 | 7622 | ChaMARCM-F000462_seg001 | Cha-Gal4 | F | 10477.58 | 0.76 |
| Cha-F-000078 | 7683 | ChaMARCM-F000505_seg001 | Cha-Gal4 | F | 10836.07 | 0.78 |
| Cha-F-000089 | 7697 | ChaMARCM-F000515_seg002 | Cha-Gal4 | F | 10627.15 | 0.76 |
| VGlut-F-000587 | 7929 | DvGlutMARCM-F003767_seg001 | VGlut-Gal4 | F | 10244.92 | 0.76 |
| VGlut-F-200534 | 7940 | DvGlutMARCM-F003775_seg001 | VGlut-Gal4 | F | 10217.32 | 0.76 |
| VGlut-F-100340 | 7978 | DvGlutMARCM-F003803_seg001 | VGlut-Gal4 | F | 8289.19 | 0.69 |
| Cha-F-400031 | 8171 | ChaMARCM-F000191_seg001 | Cha-Gal4 | F | 10400.48 | 0.75 |
| VGlut-F-000562 | 8634 | DvGlutMARCM-F003687_seg001 | VGlut-Gal4 | F | 9973.24 | 0.75 |
| VGlut-F-000549 | 8781 | DvGlutMARCM-F003599_seg001 | VGlut-Gal4 | F | 9744.23 | 0.76 |
| VGlut-F-900091 | 8964 | DvGlutMARCM-F003522_seg001 | VGlut-Gal4 | F | 9931.49 | 0.75 |
| VGlut-F-200482 | 8991 | DvGlutMARCM-F003540_seg004 | VGlut-Gal4 | F | 8849.49 | 0.66 |
| VGlut-F-600609 | 9137 | DvGlutMARCM-F003429_seg001 | VGlut-Gal4 | F | 9300.98 | 0.71 |
| VGlut-F-600587 | 9265 | DvGlutMARCM-F003322_seg001 | VGlut-Gal4 | F | 10129.39 | 0.74 |
| VGlut-F-300488 | 9372 | DvGlutMARCM-F003202_seg001 | VGlut-Gal4 | F | 9482.06 | 0.74 |
| VGlut-F-300493 | 9430 | DvGlutMARCM-F003243_seg001 | VGlut-Gal4 | F | 10005.48 | 0.73 |
| Gad1-F-700035 | 9680 | GadMARCM-F000530_seg001 | Gad1-Gal4 | F | 10555.30 | 0.77 |
| VGlut-F-200073 | 10246 | DvGlutMARCM-F198_seg1 | VGlut-Gal4 | F | 10353.16 | 0.77 |
| Gad1-F-500016 | 10500 | GadMARCM-F000153_seg001 | Gad1-Gal4 | F | 10569.23 | 0.77 |
| VGlut-F-200418 | 10771 | DvGlutMARCM-F002429_seg002 | VGlut-Gal4 | F | 10133.23 | 0.71 |
| VGlut-F-000156 | 10827 | DvGlutMARCM-F561_seg1 | VGlut-Gal4 | F | 10518.06 | 0.76 |
| VGlut-F-400604 | 10845 | DvGlutMARCM-F002771_seg001 | VGlut-Gal4 | F | 9752.11 | 0.74 |
| VGlut-F-200438 | 11057 | DvGlutMARCM-F002936_seg001 | VGlut-Gal4 | F | 9845.79 | 0.74 |
| VGlut-F-600229 | 11831 | DvGlutMARCM-F002390_seg001 | VGlut-Gal4 | F | 10337.43 | 0.75 |
| VGlut-F-400538 | 11934 | DvGlutMARCM-F002469_seg001 | VGlut-Gal4 | F | 9715.56 | 0.71 |
| VGlut-F-900033 | 11942 | DvGlutMARCM-F002476_seg001 | VGlut-Gal4 | F | 10388.75 | 0.77 |
| VGlut-F-300421 | 12046 | DvGlutMARCM-F002559_seg001 | VGlut-Gal4 | F | 10188.65 | 0.77 |
| VGlut-F-100197 | 12750 | DvGlutMARCM-F1809_seg1 | VGlut-Gal4 | F | 9880.95 | 0.75 |
| TH-F-000048 | 13658 | THMARCM-671F_seg | TH-Gal4 | F | 10333.13 | 0.48 |
| VGlut-F-100082 | 14325 | DvGlutMARCM-F1010_seg1 | VGlut-Gal4 | F | 9210.88 | 0.70 |
| VGlut-F-100093 | 14374 | DvGlutMARCM-F1054_seg1 | VGlut-Gal4 | F | 10081.50 | 0.73 |
| VGlut-F-400232 | 14546 | DvGlutMARCM-F814_seg1 | VGlut-Gal4 | F | 10146.62 | 0.76 |
| VGlut-F-100064 | 14570 | DvGlutMARCM-F835_seg1 | VGlut-Gal4 | F | 9402.14 | 0.71 |
| VGlut-F-300177 | 14575 | DvGlutMARCM-F840_seg1 | VGlut-Gal4 | F | 10421.67 | 0.77 |
| VGlut-F-000310 | 14787 | DvGlutMARCM-F1137_seg1 | VGlut-Gal4 | F | 10704.27 | 0.74 |
| VGlut-F-000312 | 14806 | DvGlutMARCM-F1153_seg1 | VGlut-Gal4 | F | 10345.96 | 0.74 |
| VGlut-F-400339 | 14875 | DvGlutMARCM-F1219_seg1 | VGlut-Gal4 | F | 10566.60 | 0.77 |
| VGlut-F-300259 | 15137 | DvGlutMARCM-F1470_seg1 | VGlut-Gal4 | F | 9647.42 | 0.71 |
| VGlut-F-000374 | 15256 | DvGlutMARCM-F1576_seg1 | VGlut-Gal4 | F | 10333.72 | 0.74 |
| VGlut-F-000379 | 15264 | DvGlutMARCM-F1584_seg1 | VGlut-Gal4 | F | 10127.71 | 0.73 |
| VGlut-F-200321 | 15376 | DvGlutMARCM-F1731_seg1 | VGlut-Gal4 | F | 10117.74 | 0.77 |
| VGlut-F-000180 | 15451 | DvGlutMARCM-F609_seg1 | VGlut-Gal4 | F | 10548.98 | 0.75 |
| VGlut-F-100000 | 15587 | DvGlutMARCM-F003_seg1 | VGlut-Gal4 | F | 10478.45 | 0.77 |
| VGlut-F-200001 | 15589 | DvGlutMARCM-F005_seg2 | VGlut-Gal4 | F | 9659.65 | 0.68 |
| VGlut-F-200006 | 15599 | DvGlutMARCM-F015_seg1 | VGlut-Gal4 | F | 9336.30 | 0.73 |
| VGlut-F-000007 | 15625 | DvGlutMARCM-F041-X2_seg1 | VGlut-Gal4 | F | 6842.87 | 0.51 |
| VGlut-F-300012 | 15685 | DvGlutMARCM-F100_seg1 | VGlut-Gal4 | F | 9992.95 | 0.72 |
| VGlut-F-000018 | 15744 | DvGlutMARCM-F154_seg1 | VGlut-Gal4 | F | 10197.00 | 0.76 |
| VGlut-F-300067 | 15959 | DvGlutMARCM-F351-X2_seg2 | VGlut-Gal4 | F | 9684.15 | 0.70 |
| VGlut-F-000108 | 16099 | DvGlutMARCM-F483_seg1 | VGlut-Gal4 | F | 9542.83 | 0.70 |
| VGlut-F-000142 | 16162 | DvGlutMARCM-F545_seg1 | VGlut-Gal4 | F | 10129.33 | 0.76 |
| VGlut-F-000165 | 16193 | DvGlutMARCM-F575_seg1 | VGlut-Gal4 | F | 9631.94 | 0.75 |
| VGlut-F-200106 | 16200 | DvGlutMARCM-F581_seg1 | VGlut-Gal4 | F | 9507.29 | 0.72 |