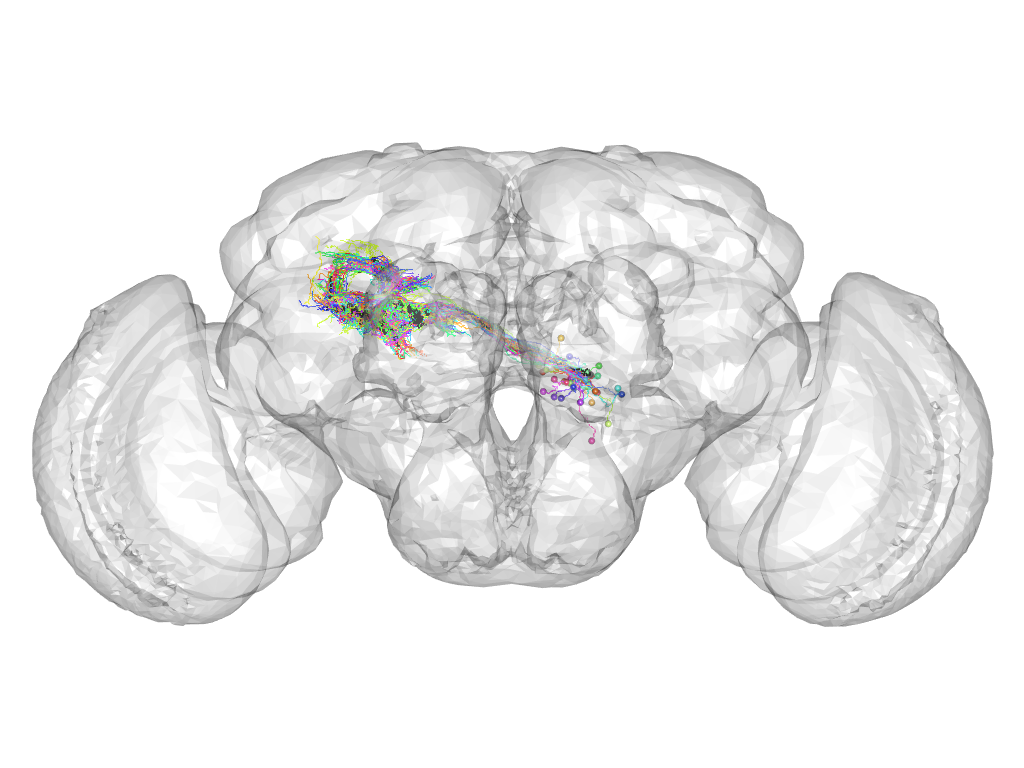
Cluster 184
Part of supercluster XIII
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (Trh-M-700031), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 166 | Trh-M-100028 | XIII | 0.72 |
| 82 | Trh-M-500117 | XIII | 0.71 |
| 798 | Trh-F-600079 | XIII | 0.58 |
| 179 | Trh-M-100061 | VI | 0.52 |
| 268 | Cha-F-200340 | X | 0.27 |
Cluster composition
This cluster has 50 neurons. The exemplar of this cluster (Trh-M-700031) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| Trh-M-700031 | 3039 | TPHMARCM-992M_seg1 | Trh-Gal4 | M | 40454.54 | 1.00 |
| fru-M-100091 | 1529 | FruMARCM-M001069_seg002 | fru-Gal4 | M | 28334.90 | 0.73 |
| Trh-M-900032 | 2039 | TPHMARCM-M001549_seg003 | Trh-Gal4 | M | 27860.56 | 0.73 |
| Trh-M-900039 | 2049 | TPHMARCM-M001554_seg003 | Trh-Gal4 | M | 29225.47 | 0.73 |
| Trh-M-900040 | 2050 | TPHMARCM-M001555_seg001 | Trh-Gal4 | M | 29348.80 | 0.74 |
| Trh-M-600087 | 2126 | TPHMARCM-M001619_seg001 | Trh-Gal4 | M | 28303.44 | 0.72 |
| Trh-M-600065 | 2608 | TPHMARCM-1315M_seg1 | Trh-Gal4 | M | 29005.94 | 0.71 |
| Trh-M-500119 | 2705 | TPHMARCM-M001469_seg001 | Trh-Gal4 | M | 27741.48 | 0.70 |
| Trh-M-500151 | 2756 | TPHMARCM-M001523_seg001 | Trh-Gal4 | M | 28005.25 | 0.74 |
| Trh-M-600056 | 2786 | TPHMARCM-1010M_seg1 | Trh-Gal4 | M | 29713.66 | 0.73 |
| Trh-M-700060 | 2809 | TPHMARCM-1039M_seg1 | Trh-Gal4 | M | 29790.60 | 0.74 |
| Trh-M-400065 | 2833 | TPHMARCM-1060M_seg2 | Trh-Gal4 | M | 12910.52 | 0.37 |
| Trh-M-800006 | 2935 | TPHMARCM-870M_seg1 | Trh-Gal4 | M | 28576.90 | 0.71 |
| Trh-M-900020 | 2957 | TPHMARCM-898M_seg1 | Trh-Gal4 | M | 28522.17 | 0.71 |
| Trh-M-600038 | 2992 | TPHMARCM-935M_seg1 | Trh-Gal4 | M | 29599.40 | 0.75 |
| Trh-M-600040 | 2994 | TPHMARCM-937M_seg1 | Trh-Gal4 | M | 28188.09 | 0.73 |
| Trh-M-600042 | 2999 | TPHMARCM-942M_seg1 | Trh-Gal4 | M | 28951.33 | 0.75 |
| Trh-M-600051 | 3021 | TPHMARCM-973M_seg1 | Trh-Gal4 | M | 27097.71 | 0.71 |
| Trh-M-600052 | 3022 | TPHMARCM-974M_seg1 | Trh-Gal4 | M | 19783.49 | 0.58 |
| Trh-M-700023 | 3026 | TPHMARCM-978M_seg1 | Trh-Gal4 | M | 26347.33 | 0.69 |
| Trh-M-000004 | 3399 | TPHMARCM-085M_seg1 | Trh-Gal4 | M | 28271.33 | 0.72 |
| Trh-M-000006 | 3402 | TPHMARCM-088M_seg1 | Trh-Gal4 | M | 29920.51 | 0.73 |
| Trh-M-500001 | 3433 | TPHMARCM-180M_seg1 | Trh-Gal4 | M | 28475.75 | 0.71 |
| Trh-M-000017 | 3480 | TPHMARCM-351M_seg1 | Trh-Gal4 | M | 28371.02 | 0.72 |
| Trh-M-500012 | 3486 | TPHMARCM-359M_seg1 | Trh-Gal4 | M | 28578.72 | 0.71 |
| Trh-M-600006 | 3547 | TPHMARCM-462M_seg1 | Trh-Gal4 | M | 28576.96 | 0.71 |
| Trh-M-500038 | 3627 | TPHMARCM-601M_seg1 | Trh-Gal4 | M | 29869.92 | 0.76 |
| Trh-F-600021 | 5993 | TPHMARCM-570F_seg2 | Trh-Gal4 | F | 26758.70 | 0.71 |
| Trh-F-600061 | 10438 | TPHMARCM-874F_seg2 | Trh-Gal4 | F | 27969.04 | 0.73 |
| Trh-F-600078 | 12419 | TPHMARCM-1211F_seg2 | Trh-Gal4 | F | 24424.55 | 0.68 |
| Trh-F-700059 | 12424 | TPHMARCM-1216F_seg1 | Trh-Gal4 | F | 26168.09 | 0.69 |
| Trh-F-600094 | 12447 | TPHMARCM-1242F_seg1 | Trh-Gal4 | F | 27659.17 | 0.73 |
| Trh-F-500155 | 12499 | TPHMARCM-1283F_seg2 | Trh-Gal4 | F | 27519.24 | 0.69 |
| Trh-F-600068 | 12730 | TPHMARCM-930F_seg3 | Trh-Gal4 | F | 27850.65 | 0.70 |
| Trh-F-900016 | 13259 | TPHMARCM-747F_seg1 | Trh-Gal4 | F | 29549.07 | 0.74 |
| Trh-F-400078 | 13270 | TPHMARCM-757F_seg2 | Trh-Gal4 | F | 27821.21 | 0.70 |
| Trh-F-800003 | 13273 | TPHMARCM-759F_seg1 | Trh-Gal4 | F | 28004.98 | 0.71 |
| Trh-F-900021 | 13282 | TPHMARCM-776F_seg1 | Trh-Gal4 | F | 29296.98 | 0.73 |
| Trh-F-600039 | 13283 | TPHMARCM-777F_seg1 | Trh-Gal4 | F | 27414.93 | 0.73 |
| Trh-F-600003 | 13898 | TPHMARCM-162F_seg2 | Trh-Gal4 | F | 28311.81 | 0.73 |
| Trh-F-600005 | 13900 | TPHMARCM-163F_seg2 | Trh-Gal4 | F | 28186.61 | 0.73 |
| Trh-F-900007 | 13926 | TPHMARCM-216F_seg2 | Trh-Gal4 | F | 23770.60 | 0.65 |
| Trh-F-500038 | 14085 | TPHMARCM-434F_seg1 | Trh-Gal4 | F | 28963.18 | 0.74 |
| Trh-F-500048 | 14136 | TPHMARCM-526F_seg1 | Trh-Gal4 | F | 29097.42 | 0.74 |
| Trh-F-600026 | 14159 | TPHMARCM-587F_seg1 | Trh-Gal4 | F | 27539.15 | 0.71 |
| Trh-F-300081 | 14166 | TPHMARCM-593F_seg2 | Trh-Gal4 | F | 24446.17 | 0.64 |
| Trh-F-500060 | 14196 | TPHMARCM-637F_seg2 | Trh-Gal4 | F | 27884.04 | 0.72 |
| Trh-F-700012 | 14224 | TPHMARCM-678F_seg1 | Trh-Gal4 | F | 29041.00 | 0.74 |
| Trh-F-500075 | 14238 | TPHMARCM-693F_seg1 | Trh-Gal4 | F | 28389.49 | 0.74 |
| Trh-F-500103 | 14259 | TPHMARCM-803F_seg1 | Trh-Gal4 | F | 28481.77 | 0.74 |