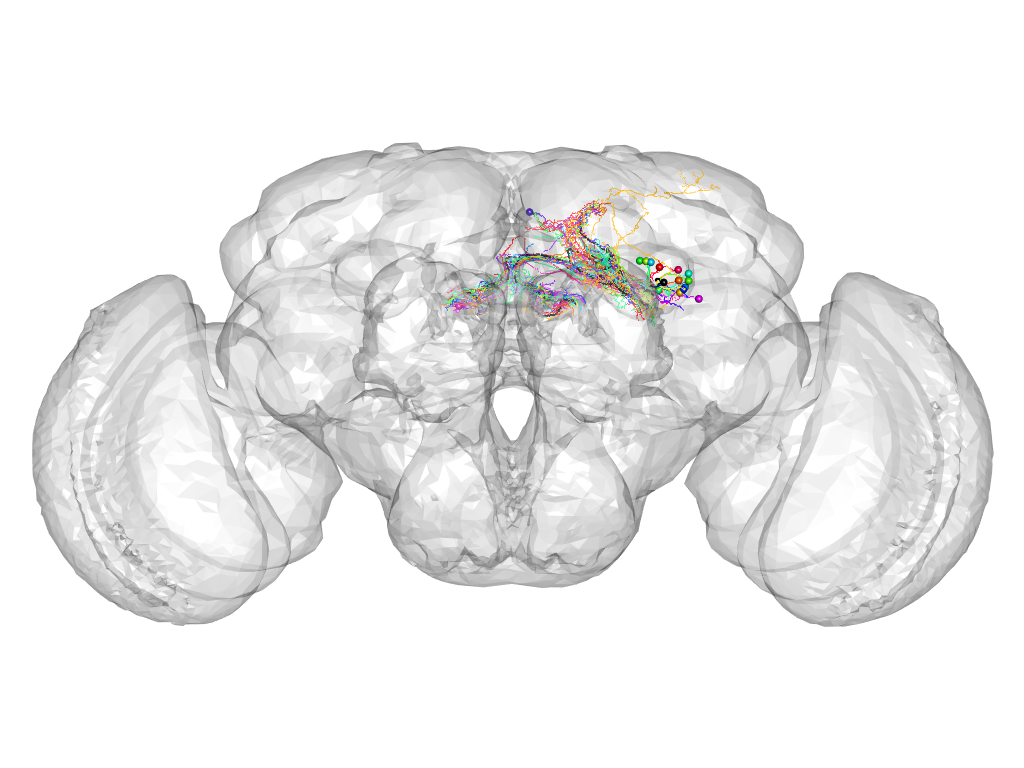
Cluster 893
Part of supercluster XIV
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (Trh-F-300061), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 213 | Trh-M-400039 | XIV | 0.70 |
| 191 | TH-M-200018 | XIV | 0.64 |
| 204 | Trh-M-100025 | XIV | 0.61 |
| 1028 | VGlut-F-000067 | XIV | 0.61 |
| 387 | Tdc2-F-000040 | XIV | 0.60 |
| 667 | TH-F-300071 | XIV | 0.55 |
Cluster composition
This cluster has 18 neurons. The exemplar of this cluster (Trh-F-300061) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| Trh-F-300061 | 14105 | TPHMARCM-464F_seg1 | Trh-Gal4 | F | 11935.91 | 1.00 |
| Trh-M-500105 | 926 | TPHMARCM-M001450_seg001 | Trh-Gal4 | M | 8062.62 | 0.63 |
| Trh-M-500178 | 2083 | TPHMARCM-M001584_seg001 | Trh-Gal4 | M | 8359.73 | 0.69 |
| Trh-M-600111 | 2217 | TPHMARCM-M001708_seg001 | Trh-Gal4 | M | 7145.57 | 0.45 |
| Trh-M-500198 | 2224 | TPHMARCM-M001714_seg001 | Trh-Gal4 | M | 8123.06 | 0.69 |
| Trh-M-500103 | 2682 | TPHMARCM-M001448_seg001 | Trh-Gal4 | M | 8213.93 | 0.72 |
| Trh-M-600078 | 2730 | TPHMARCM-M001495_seg001 | Trh-Gal4 | M | 8620.17 | 0.73 |
| Trh-M-500137 | 2734 | TPHMARCM-M001502_seg001 | Trh-Gal4 | M | 8035.23 | 0.67 |
| Cha-F-500061 | 8178 | ChaMARCM-F000195_seg001 | Cha-Gal4 | F | 6926.59 | 0.60 |
| Gad1-F-200088 | 11334 | GadMARCM-F000302_seg001 | Gad1-Gal4 | F | 6703.00 | 0.63 |
| Trh-F-600073 | 12399 | TPHMARCM-1196F_seg1 | Trh-Gal4 | F | 8614.46 | 0.71 |
| Trh-F-600101 | 12462 | TPHMARCM-1250F_seg1 | Trh-Gal4 | F | 7199.20 | 0.67 |
| Trh-F-700048 | 12724 | TPHMARCM-885F_seg1 | Trh-Gal4 | F | 8561.77 | 0.71 |
| Trh-F-700022 | 13244 | TPHMARCM-731F_seg1 | Trh-Gal4 | F | 8456.45 | 0.70 |
| Trh-F-500084 | 13253 | TPHMARCM-740F_seg1 | Trh-Gal4 | F | 8472.63 | 0.69 |
| Trh-F-500055 | 14181 | TPHMARCM-620F_seg1 | Trh-Gal4 | F | 8548.25 | 0.70 |
| Trh-F-400063 | 14200 | TPHMARCM-641F_seg1 | Trh-Gal4 | F | 6862.53 | 0.61 |
| Trh-F-700036 | 14272 | TPHMARCM-820F_seg1 | Trh-Gal4 | F | 8809.47 | 0.70 |