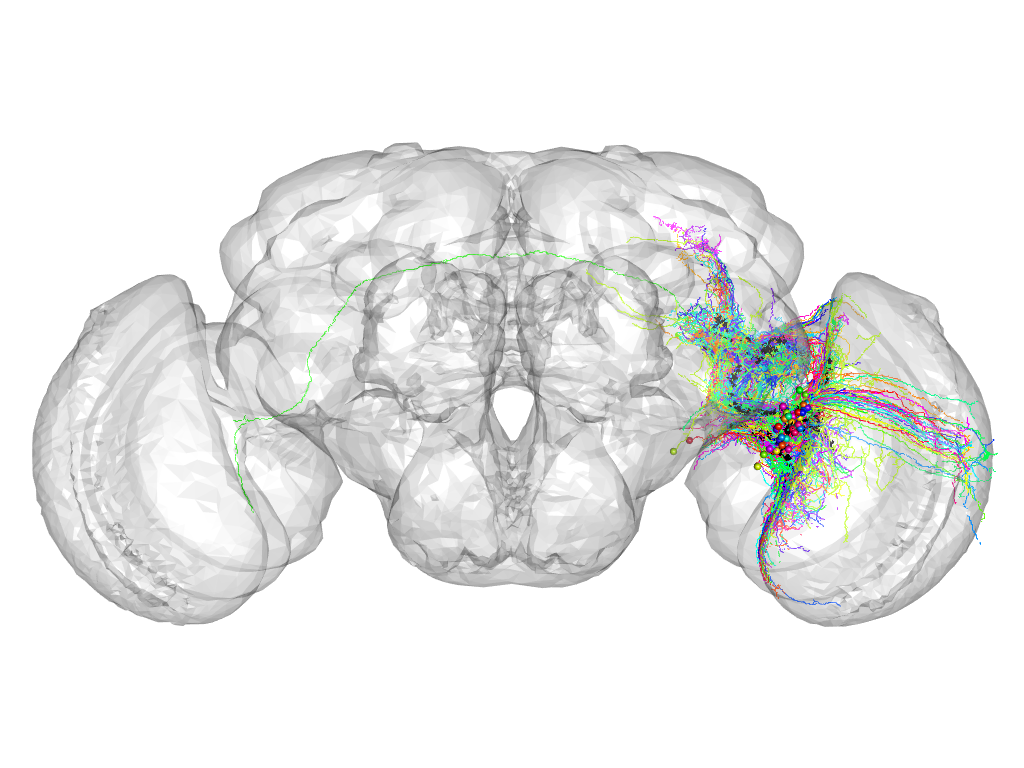
Cluster 849
Part of supercluster I
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (TH-F-300018), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 870 | TH-F-000064 | I | 0.71 |
| 874 | Trh-F-100011 | III | 0.67 |
| 989 | VGlut-F-200014 | I | 0.66 |
| 65 | fru-M-000139 | XVII | 0.57 |
| 1006 | VGlut-F-500043 | III | 0.50 |
Cluster composition
This cluster has 48 neurons. The exemplar of this cluster (TH-F-300018) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| TH-F-300018 | 13399 | THMARCM-189F_seg1 | TH-Gal4 | F | 85214.22 | 1.00 |
| Trh-M-000055 | 2063 | TPHMARCM-M001568_seg001 | Trh-Gal4 | M | 59477.05 | 0.64 |
| Trh-M-000063 | 2210 | TPHMARCM-M001701_seg001 | Trh-Gal4 | M | 52667.54 | 0.61 |
| TH-M-100073 | 3051 | THMARCM-572M_seg1 | TH-Gal4 | M | 56467.11 | 0.64 |
| TH-M-100079 | 3058 | THMARCM-614M_seg | TH-Gal4 | M | 59906.17 | 0.67 |
| TH-M-900000 | 3064 | THMARCM-637M_seg1 | TH-Gal4 | M | 59105.04 | 0.64 |
| TH-M-700000 | 3075 | THMARCM-021M_seg1 | TH-Gal4 | M | 57468.41 | 0.60 |
| TH-M-300002 | 3083 | THMARCM-054M_seg1 | TH-Gal4 | M | 59111.84 | 0.61 |
| Trh-M-100049 | 3578 | TPHMARCM-508M_seg1 | Trh-Gal4 | M | 60819.93 | 0.68 |
| fru-F-200176 | 3811 | FruMARCM-F002567_seg001 | fru-Gal4 | F | 59735.08 | 0.64 |
| Cha-F-300333 | 3888 | ChaMARCM-F001550_seg001 | Cha-Gal4 | F | 58295.59 | 0.64 |
| Cha-F-300310 | 4439 | ChaMARCM-F001482_seg001 | Cha-Gal4 | F | 56512.65 | 0.50 |
| Cha-F-200247 | 5154 | ChaMARCM-F001200_seg001 | Cha-Gal4 | F | 59347.21 | 0.60 |
| VGlut-F-400517 | 11806 | DvGlutMARCM-F002371_seg001 | VGlut-Gal4 | F | 47006.69 | 0.60 |
| TH-F-200107 | 12337 | THMARCM-683F_seg1 | TH-Gal4 | F | 58986.23 | 0.65 |
| Trh-F-100090 | 12604 | TPHMARCM-F001399_seg001 | Trh-Gal4 | F | 57086.26 | 0.64 |
| VGlut-F-300379 | 12978 | DvGlutMARCM-F2012-montage_seg1 | VGlut-Gal4 | F | 34669.32 | 0.19 |
| TH-F-000001 | 13315 | THMARCM-031F_seg2 | TH-Gal4 | F | 53577.64 | 0.61 |
| TH-F-000002 | 13316 | THMARCM-033F_seg1 | TH-Gal4 | F | 57612.83 | 0.68 |
| TH-F-100004 | 13320 | THMARCM-038F_seg1 | TH-Gal4 | F | 63300.90 | 0.66 |
| TH-F-300000 | 13321 | THMARCM-041F_seg1 | TH-Gal4 | F | 59471.42 | 0.62 |
| TH-F-100006 | 13323 | THMARCM-043F_seg1 | TH-Gal4 | F | 60794.98 | 0.68 |
| TH-F-400000 | 13329 | THMARCM-052F_seg1 | TH-Gal4 | F | 58451.96 | 0.60 |
| TH-F-300001 | 13333 | THMARCM-061F_seg1 | TH-Gal4 | F | 57430.75 | 0.62 |
| TH-F-300005 | 13346 | THMARCM-100F_seg1 | TH-Gal4 | F | 57424.13 | 0.61 |
| TH-F-400011 | 13393 | THMARCM-180F_seg1 | TH-Gal4 | F | 59159.04 | 0.67 |
| TH-F-700037 | 13394 | THMARCM-185F_seg1 | TH-Gal4 | F | 57951.26 | 0.63 |
| TH-F-700039 | 13395 | THMARCM-186F_seg2 | TH-Gal4 | F | 57601.39 | 0.63 |
| TH-F-300017 | 13398 | THMARCM-188F_seg1 | TH-Gal4 | F | 54539.21 | 0.58 |
| TH-F-100017 | 13409 | THMARCM-210F_seg1 | TH-Gal4 | F | 59694.84 | 0.71 |
| TH-F-300026 | 13418 | THMARCM-224F_seg2 | TH-Gal4 | F | 58687.03 | 0.64 |
| TH-F-300027 | 13419 | THMARCM-226F_seg1 | TH-Gal4 | F | 56425.33 | 0.64 |
| TH-F-300028 | 13421 | THMARCM-229F_seg1 | TH-Gal4 | F | 53690.93 | 0.64 |
| TH-F-300030 | 13423 | THMARCM-232F_seg1 | TH-Gal4 | F | 56703.32 | 0.61 |
| TH-F-500000 | 13629 | THMARCM-611F_seg | TH-Gal4 | F | 60580.14 | 0.68 |
| TH-F-000032 | 13631 | THMARCM-613F_seg | TH-Gal4 | F | 58082.74 | 0.64 |
| TH-F-000038 | 13641 | THMARCM-644F_seg | TH-Gal4 | F | 61317.10 | 0.71 |
| TH-F-000039 | 13642 | THMARCM-645F_seg | TH-Gal4 | F | 54404.93 | 0.58 |
| TH-F-000042 | 13647 | THMARCM-650F-X2_seg2 | TH-Gal4 | F | 56701.65 | 0.65 |
| Trh-F-100005 | 13788 | TPHMARCM-006F_seg1 | Trh-Gal4 | F | 57258.80 | 0.60 |
| Trh-F-100013 | 13803 | TPHMARCM-021F_seg1 | Trh-Gal4 | F | 60567.65 | 0.60 |
| Trh-F-100019 | 13812 | TPHMARCM-031F_seg1 | Trh-Gal4 | F | 58956.95 | 0.65 |
| Trh-F-200020 | 13942 | TPHMARCM-234F_seg1 | Trh-Gal4 | F | 56853.76 | 0.60 |
| Trh-F-100039 | 14036 | TPHMARCM-326F_seg1 | Trh-Gal4 | F | 57515.61 | 0.65 |
| Trh-F-100048 | 14059 | TPHMARCM-386F_seg1 | Trh-Gal4 | F | 58166.83 | 0.61 |
| Trh-F-100055 | 14161 | TPHMARCM-588F_seg1 | Trh-Gal4 | F | 55275.13 | 0.64 |
| VGlut-F-200252 | 14920 | DvGlutMARCM-F1262_seg1 | VGlut-Gal4 | F | 59950.83 | 0.67 |
| VGlut-F-900024 | 14977 | DvGlutMARCM-F1316_seg1 | VGlut-Gal4 | F | 48661.57 | 0.61 |