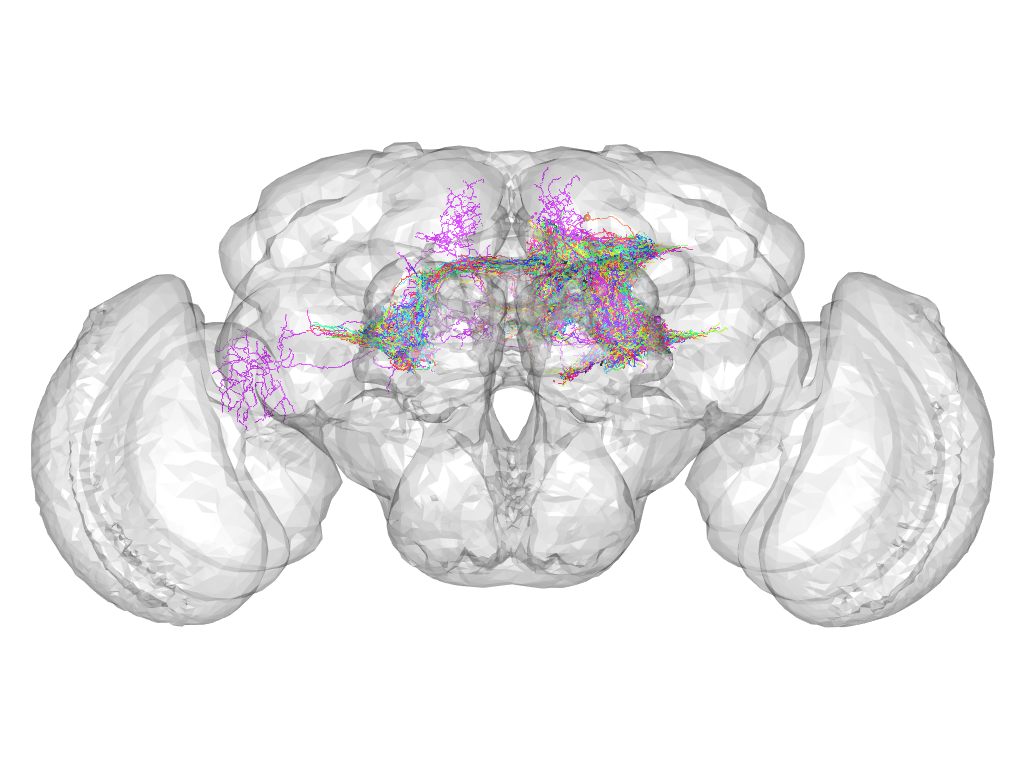
Cluster 836
Part of supercluster VI
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (VGlut-F-100232), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 907 | VGlut-F-000294 | VI | 0.58 |
| 353 | Cha-F-200264 | VI | 0.58 |
| 686 | VGlut-F-900065 | VII | 0.52 |
| 318 | fru-F-600121 | VI | 0.47 |
| 755 | VGlut-F-200398 | VI | 0.45 |
Cluster composition
This cluster has 92 neurons. The exemplar of this cluster (VGlut-F-100232) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| VGlut-F-100232 | 13130 | DvGlutMARCM-F2157_seg1 | VGlut-Gal4 | F | 28256.68 | 1.00 |
| fru-M-200279 | 116 | FruMARCM-M002314_seg001 | fru-Gal4 | M | 19828.46 | 0.69 |
| fru-M-100174 | 712 | FruMARCM-M001916_seg001 | fru-Gal4 | M | 19218.77 | 0.68 |
| fru-M-200139 | 1335 | FruMARCM-M001092_seg001 | fru-Gal4 | M | 19244.38 | 0.69 |
| fru-M-200133 | 1547 | FruMARCM-M001080_seg002 | fru-Gal4 | M | 19899.03 | 0.70 |
| Cha-F-100313 | 4352 | ChaMARCM-F001410_seg001 | Cha-Gal4 | F | 14686.26 | 0.59 |
| Cha-F-400203 | 5633 | ChaMARCM-F000912_seg001 | Cha-Gal4 | F | 20421.10 | 0.72 |
| VGlut-F-700556 | 5727 | DvGlutMARCM-F004435_seg001 | VGlut-Gal4 | F | 19648.01 | 0.71 |
| VGlut-F-700607 | 5948 | DvGlutMARCM-F004602_seg001 | VGlut-Gal4 | F | 19356.20 | 0.69 |
| VGlut-F-400926 | 5949 | DvGlutMARCM-F004603_seg001 | VGlut-Gal4 | F | 20201.42 | 0.72 |
| VGlut-F-400611 | 5971 | DvGlutMARCM-F002818_seg002 | VGlut-Gal4 | F | 19175.73 | 0.69 |
| VGlut-F-700080 | 6032 | DvGlutMARCM-F002593_seg001 | VGlut-Gal4 | F | 19101.04 | 0.70 |
| VGlut-F-600657 | 6071 | DvGlutMARCM-F003533_seg001 | VGlut-Gal4 | F | 19728.55 | 0.69 |
| VGlut-F-100365 | 6414 | DvGlutMARCM-F004206_seg001 | VGlut-Gal4 | F | 19949.94 | 0.70 |
| VGlut-F-600782 | 6455 | DvGlutMARCM-F004240_seg001 | VGlut-Gal4 | F | 19306.05 | 0.68 |
| VGlut-F-300617 | 6473 | DvGlutMARCM-F004255_seg003 | VGlut-Gal4 | F | 19883.52 | 0.71 |
| VGlut-F-500794 | 6488 | DvGlutMARCM-F004266_seg001 | VGlut-Gal4 | F | 20184.37 | 0.69 |
| VGlut-F-500816 | 6599 | DvGlutMARCM-F004357_seg002 | VGlut-Gal4 | F | 20285.65 | 0.72 |
| VGlut-F-900132 | 6667 | DvGlutMARCM-F004406_seg002 | VGlut-Gal4 | F | 20391.67 | 0.71 |
| VGlut-F-100352 | 6676 | DvGlutMARCM-F003898_seg001 | VGlut-Gal4 | F | 19708.53 | 0.70 |
| VGlut-F-300561 | 6679 | DvGlutMARCM-F003901_seg001 | VGlut-Gal4 | F | 19799.32 | 0.70 |
| VGlut-F-200547 | 6743 | DvGlutMARCM-F003948_seg001 | VGlut-Gal4 | F | 19502.00 | 0.69 |
| VGlut-F-400819 | 6757 | DvGlutMARCM-F003961_seg001 | VGlut-Gal4 | F | 19279.49 | 0.67 |
| VGlut-F-500745 | 6758 | DvGlutMARCM-F003962_seg001 | VGlut-Gal4 | F | 19544.43 | 0.69 |
| VGlut-F-500749 | 6767 | DvGlutMARCM-F003969_seg002 | VGlut-Gal4 | F | 18886.17 | 0.68 |
| VGlut-F-100354 | 6801 | DvGlutMARCM-F003991_seg002 | VGlut-Gal4 | F | 19872.11 | 0.71 |
| VGlut-F-500758 | 6819 | DvGlutMARCM-F004003_seg002 | VGlut-Gal4 | F | 20272.86 | 0.71 |
| VGlut-F-000621 | 6836 | DvGlutMARCM-F004019_seg001 | VGlut-Gal4 | F | 18876.38 | 0.69 |
| VGlut-F-600729 | 6843 | DvGlutMARCM-F004022_seg003 | VGlut-Gal4 | F | 19777.32 | 0.72 |
| VGlut-F-300578 | 6897 | DvGlutMARCM-F004053_seg001 | VGlut-Gal4 | F | 20029.94 | 0.69 |
| VGlut-F-300580 | 6900 | DvGlutMARCM-F004057_seg001 | VGlut-Gal4 | F | 19285.15 | 0.68 |
| VGlut-F-000635 | 6949 | DvGlutMARCM-F004103_seg001 | VGlut-Gal4 | F | 19584.31 | 0.71 |
| VGlut-F-400866 | 6965 | DvGlutMARCM-F004113_seg002 | VGlut-Gal4 | F | 20388.60 | 0.71 |
| VGlut-F-000640 | 6968 | DvGlutMARCM-F004115_seg002 | VGlut-Gal4 | F | 19262.27 | 0.71 |
| VGlut-F-300593 | 6991 | DvGlutMARCM-F004135_seg003 | VGlut-Gal4 | F | 18008.67 | 0.68 |
| VGlut-F-400868 | 6993 | DvGlutMARCM-F004136_seg002 | VGlut-Gal4 | F | 19680.84 | 0.70 |
| VGlut-F-400873 | 6999 | DvGlutMARCM-F004141_seg001 | VGlut-Gal4 | F | 20138.63 | 0.72 |
| VGlut-F-700507 | 7049 | DvGlutMARCM-F004179_seg001 | VGlut-Gal4 | F | 20094.58 | 0.72 |
| VGlut-F-200531 | 7936 | DvGlutMARCM-F003771_seg001 | VGlut-Gal4 | F | 19458.80 | 0.69 |
| VGlut-F-600513 | 8545 | DvGlutMARCM-F003117_seg002 | VGlut-Gal4 | F | 19264.39 | 0.69 |
| VGlut-F-100318 | 8586 | DvGlutMARCM-F003652_seg001 | VGlut-Gal4 | F | 19665.91 | 0.71 |
| VGlut-F-000573 | 8666 | DvGlutMARCM-F003713_seg002 | VGlut-Gal4 | F | 20433.46 | 0.72 |
| VGlut-F-300530 | 8705 | DvGlutMARCM-F003739_seg001 | VGlut-Gal4 | F | 19379.99 | 0.69 |
| VGlut-F-000533 | 8727 | DvGlutMARCM-F003559_seg001 | VGlut-Gal4 | F | 18806.24 | 0.68 |
| VGlut-F-500707 | 8798 | DvGlutMARCM-F003611_seg002 | VGlut-Gal4 | F | 19826.35 | 0.72 |
| VGlut-F-100311 | 8842 | DvGlutMARCM-F003637_seg002 | VGlut-Gal4 | F | 17590.55 | 0.65 |
| VGlut-F-800220 | 8847 | DvGlutMARCM-F003640_seg001 | VGlut-Gal4 | F | 19830.01 | 0.68 |
| VGlut-F-900092 | 8965 | DvGlutMARCM-F003523_seg001 | VGlut-Gal4 | F | 18969.03 | 0.70 |
| VGlut-F-600653 | 8968 | DvGlutMARCM-F003527_seg001 | VGlut-Gal4 | F | 19921.10 | 0.72 |
| VGlut-F-600656 | 8972 | DvGlutMARCM-F003532_seg001 | VGlut-Gal4 | F | 19691.11 | 0.70 |
| VGlut-F-600603 | 9064 | DvGlutMARCM-F003382_seg001 | VGlut-Gal4 | F | 19783.28 | 0.70 |
| VGlut-F-200464 | 9189 | DvGlutMARCM-F003257_seg001 | VGlut-Gal4 | F | 19751.59 | 0.71 |
| VGlut-F-700318 | 9247 | DvGlutMARCM-F003307_seg001 | VGlut-Gal4 | F | 20644.83 | 0.73 |
| VGlut-F-200292 | 10233 | DvGlutMARCM-F1593_seg1 | VGlut-Gal4 | F | 20064.53 | 0.73 |
| VGlut-F-400621 | 10738 | DvGlutMARCM-F002852_seg001 | VGlut-Gal4 | F | 19385.47 | 0.69 |
| VGlut-F-400606 | 10885 | DvGlutMARCM-F002791_seg002 | VGlut-Gal4 | F | 19067.71 | 0.70 |
| VGlut-F-700200 | 10936 | DvGlutMARCM-F002829_seg002 | VGlut-Gal4 | F | 19153.67 | 0.70 |
| VGlut-F-800082 | 10961 | DvGlutMARCM-F002856_seg001 | VGlut-Gal4 | F | 20103.62 | 0.71 |
| VGlut-F-800084 | 10963 | DvGlutMARCM-F002857_seg001 | VGlut-Gal4 | F | 19684.83 | 0.70 |
| VGlut-F-900070 | 11125 | DvGlutMARCM-F002990_seg001 | VGlut-Gal4 | F | 19312.00 | 0.70 |
| VGlut-F-300439 | 11646 | DvGlutMARCM-F002221_seg001 | VGlut-Gal4 | F | 18680.72 | 0.68 |
| VGlut-F-600501 | 11653 | DvGlutMARCM-F002227_seg001 | VGlut-Gal4 | F | 20197.31 | 0.71 |
| VGlut-F-400541 | 11936 | DvGlutMARCM-F002471_seg002 | VGlut-Gal4 | F | 19759.14 | 0.70 |
| VGlut-F-900031 | 11940 | DvGlutMARCM-F002475_seg001 | VGlut-Gal4 | F | 18630.71 | 0.68 |
| VGlut-F-500457 | 12213 | DvGlutMARCM-F002681_seg001 | VGlut-Gal4 | F | 18889.99 | 0.68 |
| VGlut-F-400567 | 12226 | DvGlutMARCM-F002691_seg001 | VGlut-Gal4 | F | 18844.70 | 0.67 |
| VGlut-F-500464 | 12230 | DvGlutMARCM-F002694_seg001 | VGlut-Gal4 | F | 18976.01 | 0.69 |
| VGlut-F-500470 | 12265 | DvGlutMARCM-F002722_seg002 | VGlut-Gal4 | F | 19968.10 | 0.71 |
| VGlut-F-400588 | 12294 | DvGlutMARCM-F002748_seg002 | VGlut-Gal4 | F | 18915.45 | 0.70 |
| VGlut-F-400592 | 12303 | DvGlutMARCM-F002763_seg002 | VGlut-Gal4 | F | 19452.72 | 0.65 |
| VGlut-F-200449 | 12312 | DvGlutMARCM-F2204_seg1 | VGlut-Gal4 | F | 19782.72 | 0.70 |
| VGlut-F-200060 | 12364 | DvGlutMARCM-F113_seg1 | VGlut-Gal4 | F | 13620.50 | 0.61 |
| VGlut-F-500271 | 12789 | DvGlutMARCM-F1837_seg1 | VGlut-Gal4 | F | 18828.35 | 0.66 |
| VGlut-F-200357 | 12974 | DvGlutMARCM-F2009_seg1 | VGlut-Gal4 | F | 18746.74 | 0.68 |
| VGlut-F-000434 | 12977 | DvGlutMARCM-F2011-montage_seg1 | VGlut-Gal4 | F | 12181.67 | 0.20 |
| VGlut-F-000441 | 13040 | DvGlutMARCM-F2064_seg1 | VGlut-Gal4 | F | 19248.84 | 0.70 |
| VGlut-F-300386 | 13042 | DvGlutMARCM-F2066_seg1 | VGlut-Gal4 | F | 18830.23 | 0.69 |
| VGlut-F-700043 | 13069 | DvGlutMARCM-F2092_seg1 | VGlut-Gal4 | F | 19423.10 | 0.71 |
| VGlut-F-400308 | 14389 | DvGlutMARCM-F1069_seg1 | VGlut-Gal4 | F | 19746.57 | 0.69 |
| VGlut-F-000278 | 14391 | DvGlutMARCM-F1071_seg1 | VGlut-Gal4 | F | 18434.00 | 0.65 |
| VGlut-F-300171 | 14560 | DvGlutMARCM-F825_seg1 | VGlut-Gal4 | F | 20346.99 | 0.72 |
| VGlut-F-300204 | 14754 | DvGlutMARCM-F1105_seg1 | VGlut-Gal4 | F | 19767.14 | 0.70 |
| VGlut-F-200217 | 14820 | DvGlutMARCM-F1164_seg1 | VGlut-Gal4 | F | 19726.03 | 0.70 |
| VGlut-F-300268 | 15159 | DvGlutMARCM-F1490_seg1 | VGlut-Gal4 | F | 19622.04 | 0.69 |
| VGlut-F-300290 | 15237 | DvGlutMARCM-F1559_seg1 | VGlut-Gal4 | F | 19349.04 | 0.70 |
| VGlut-F-300296 | 15243 | DvGlutMARCM-F1564_seg1 | VGlut-Gal4 | F | 20472.36 | 0.71 |
| VGlut-F-000181 | 15452 | DvGlutMARCM-F610_seg1 | VGlut-Gal4 | F | 20139.32 | 0.71 |
| VGlut-F-200059 | 15695 | DvGlutMARCM-F110_seg1 | VGlut-Gal4 | F | 18910.47 | 0.67 |
| VGlut-F-300015 | 15721 | DvGlutMARCM-F134_seg1 | VGlut-Gal4 | F | 19725.69 | 0.70 |
| VGlut-F-200069 | 15781 | DvGlutMARCM-F187_seg1 | VGlut-Gal4 | F | 19979.27 | 0.69 |
| VGlut-F-200070 | 15782 | DvGlutMARCM-F188_seg1 | VGlut-Gal4 | F | 19601.82 | 0.69 |
| VGlut-F-200078 | 15823 | DvGlutMARCM-F232_seg1 | VGlut-Gal4 | F | 20461.32 | 0.72 |