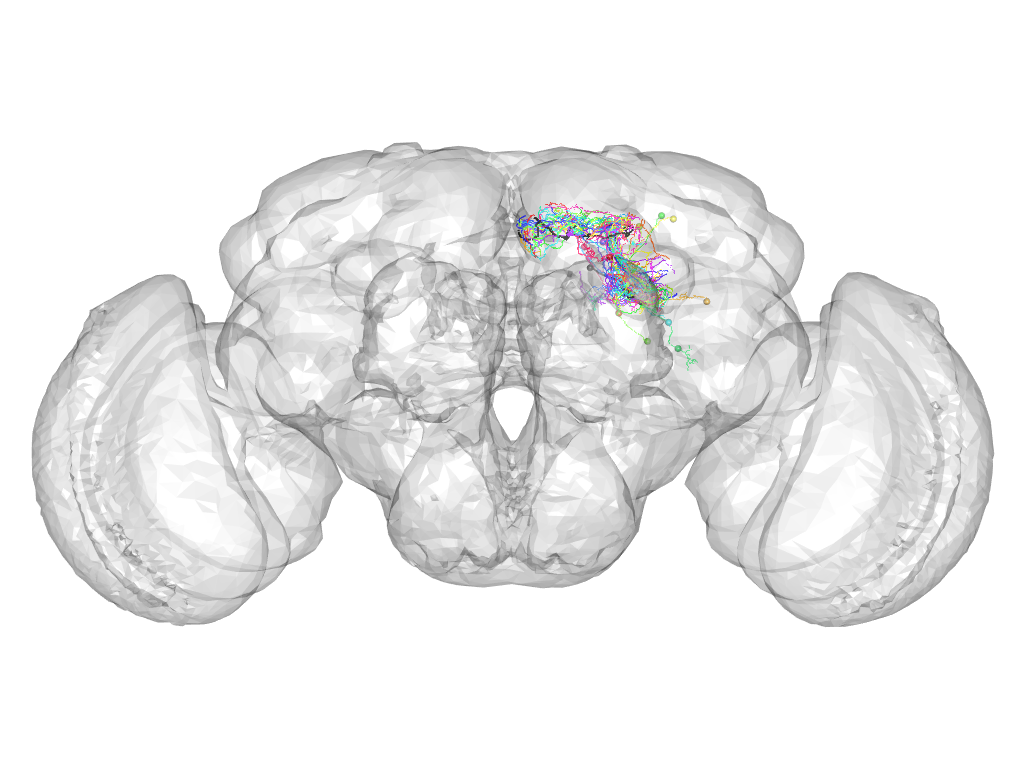
Cluster 741
Part of supercluster XV
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (fru-F-200020), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 1048 | VGlut-F-000167 | XV | 0.53 |
| 161 | fru-M-100017 | XV | 0.51 |
| 295 | fru-F-100054 | XV | 0.48 |
| 745 | Cha-F-300003 | XV | 0.44 |
| 671 | Gad1-F-300023 | XV | 0.42 |
Cluster composition
This cluster has 19 neurons. The exemplar of this cluster (fru-F-200020) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| fru-F-200020 | 11516 | FruMARCM-F000176_seg002 | fru-Gal4 | F | 6571.59 | 1.00 |
| fru-M-100163 | 533 | FruMARCM-M001652_seg001 | fru-Gal4 | M | 3135.70 | 0.52 |
| fru-M-800068 | 586 | FruMARCM-M001779_seg001 | fru-Gal4 | M | 2004.45 | 0.37 |
| fru-M-000111 | 600 | FruMARCM-M001792_seg001 | fru-Gal4 | M | 3894.95 | 0.54 |
| fru-M-300097 | 1360 | FruMARCM-M000870_seg002 | fru-Gal4 | M | 3911.43 | 0.52 |
| fru-M-000030 | 1671 | FruMARCM-M000829_seg001 | fru-Gal4 | M | 3202.24 | 0.50 |
| 5-HT1B-M-300002 | 2512 | 5HT1bMARCM-M000054_seg002 | 5-HT1B-Gal4 | M | 3618.56 | 0.56 |
| 5-HT1B-M-000005 | 2532 | 5HT1bMARCM-M000072_seg002 | 5-HT1B-Gal4 | M | 3084.70 | 0.47 |
| fru-F-100090 | 3805 | FruMARCM-F002560_seg001 | fru-Gal4 | F | 3083.12 | 0.43 |
| Gad1-F-000094 | 4106 | GadMARCM-F000692_seg003 | Gad1-Gal4 | F | 3587.65 | 0.51 |
| VGlut-F-100359 | 6939 | DvGlutMARCM-F004093_seg001 | VGlut-Gal4 | F | 3821.28 | 0.54 |
| fru-F-200065 | 7888 | FruMARCM-F000819_seg001 | fru-Gal4 | F | 3172.20 | 0.51 |
| Gad1-F-400050 | 8411 | GadMARCM-F000348_seg001 | Gad1-Gal4 | F | 3916.30 | 0.57 |
| Gad1-F-100043 | 9602 | GadMARCM-F000469_seg001 | Gad1-Gal4 | F | 3154.98 | 0.47 |
| 5-HT1B-F-300001 | 11202 | 5HT1bMARCM-F000031_seg001 | 5-HT1B-Gal4 | F | 3245.87 | 0.50 |
| Gad1-F-400026 | 11246 | GadMARCM-F000226_seg001 | Gad1-Gal4 | F | 3404.04 | 0.48 |
| VGlut-F-600286 | 11638 | DvGlutMARCM-F002523_seg005 | VGlut-Gal4 | F | 2094.00 | 0.40 |
| VGlut-F-100252 | 11692 | DvGlutMARCM-F002266_seg002 | VGlut-Gal4 | F | 2992.27 | 0.47 |
| VGlut-F-200421 | 11909 | DvGlutMARCM-F002445_seg001 | VGlut-Gal4 | F | 3202.61 | 0.50 |