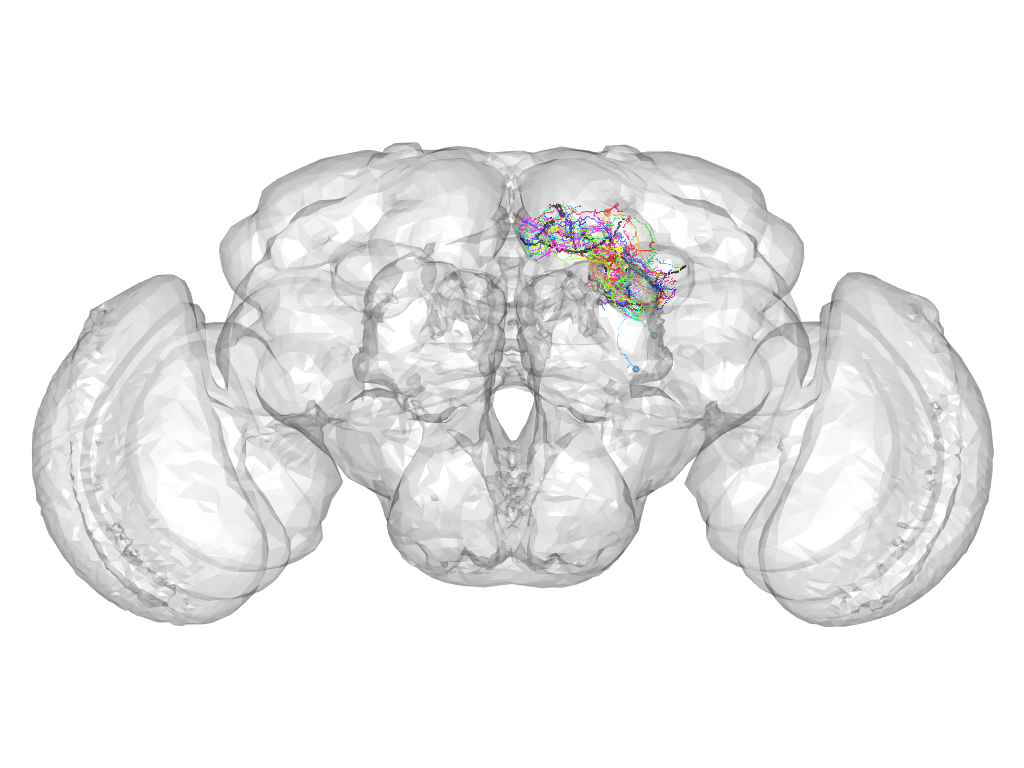
Cluster 745
Part of supercluster XV
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (Cha-F-300003), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 1048 | VGlut-F-000167 | XV | 0.50 |
| 161 | fru-M-100017 | XV | 0.46 |
| 169 | Trh-M-200075 | XV | 0.44 |
| 615 | Gad1-F-400089 | XV | 0.43 |
| 671 | Gad1-F-300023 | XV | 0.42 |
Cluster composition
This cluster has 20 neurons. The exemplar of this cluster (Cha-F-300003) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| Cha-F-300003 | 11573 | ChaMARCM-F000010_seg002 | Cha-Gal4 | F | 12083.97 | 1.00 |
| fru-M-900046 | 809 | FruMARCM-M002017_seg001 | fru-Gal4 | M | 3079.20 | 0.41 |
| fru-M-600039 | 989 | FruMARCM-M001171_seg001 | fru-Gal4 | M | 4553.79 | 0.44 |
| fru-M-100001 | 2351 | FruMARCM-M000044_seg001 | fru-Gal4 | M | 5475.82 | 0.47 |
| fru-M-300023 | 2405 | FruMARCM-M000127_seg002 | fru-Gal4 | M | 6678.45 | 0.60 |
| fru-F-000140 | 3807 | FruMARCM-F002562_seg001 | fru-Gal4 | F | 6192.58 | 0.55 |
| VGlut-F-100392 | 5724 | DvGlutMARCM-F004431_seg001 | VGlut-Gal4 | F | 6044.20 | 0.57 |
| Cha-F-300143 | 7728 | ChaMARCM-F000539_seg002 | Cha-Gal4 | F | 3995.76 | 0.39 |
| Gad1-F-700030 | 9502 | GadMARCM-F000394_seg002 | Gad1-Gal4 | F | 6570.94 | 0.60 |
| Gad1-F-700031 | 9567 | GadMARCM-F000441_seg001 | Gad1-Gal4 | F | 6073.82 | 0.56 |
| Gad1-F-200118 | 9621 | GadMARCM-F000482_seg001 | Gad1-Gal4 | F | 5564.17 | 0.54 |
| Gad1-F-200038 | 10497 | GadMARCM-F000151_seg002 | Gad1-Gal4 | F | 4780.64 | 0.52 |
| Gad1-F-200021 | 10727 | GadMARCM-F000054_seg002 | Gad1-Gal4 | F | 3221.23 | 0.32 |
| Gad1-F-200061 | 11239 | GadMARCM-F000220_seg002 | Gad1-Gal4 | F | 6423.26 | 0.61 |
| Gad1-F-000022 | 11265 | GadMARCM-F000243_seg001 | Gad1-Gal4 | F | 5422.43 | 0.51 |
| Gad1-F-800015 | 11305 | GadMARCM-F000278_seg001 | Gad1-Gal4 | F | 6833.10 | 0.58 |
| Gad1-F-400052 | 11398 | GadMARCM-F000349_seg001 | Gad1-Gal4 | F | 6338.99 | 0.56 |
| fru-F-400012 | 11469 | FruMARCM-F000082_seg001 | fru-Gal4 | F | 6477.08 | 0.55 |
| VGlut-F-100224 | 13104 | DvGlutMARCM-F2129_seg1 | VGlut-Gal4 | F | 5246.07 | 0.48 |
| VGlut-F-400306 | 14387 | DvGlutMARCM-F1067_seg1 | VGlut-Gal4 | F | 3817.26 | 0.41 |