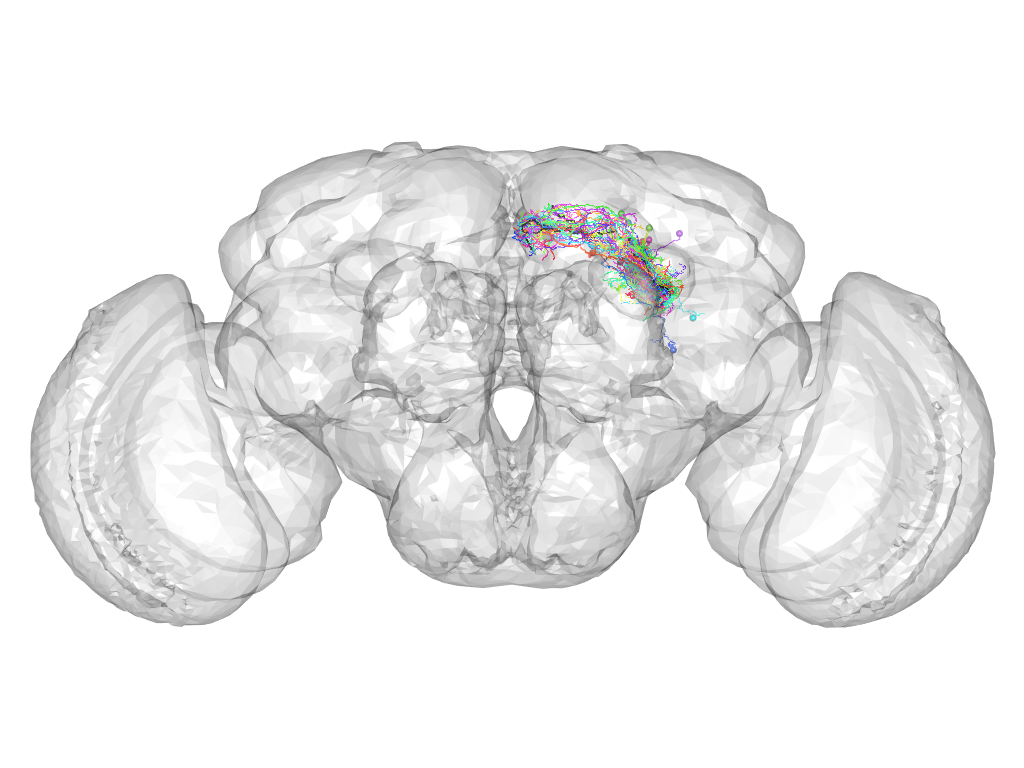
Cluster 736
Part of supercluster XV
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (fru-F-400017), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 161 | fru-M-100017 | XV | 0.53 |
| 671 | Gad1-F-300023 | XV | 0.53 |
| 1048 | VGlut-F-000167 | XV | 0.50 |
| 51 | fru-M-000088 | XV | 0.49 |
| 743 | fru-F-000002 | XV | 0.48 |
Cluster composition
This cluster has 23 neurons. The exemplar of this cluster (fru-F-400017) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| fru-F-400017 | 11472 | FruMARCM-F000085_seg001 | fru-Gal4 | F | 5204.88 | 1.00 |
| fru-M-100237 | 405 | FruMARCM-M002645_seg001 | fru-Gal4 | M | 3159.09 | 0.56 |
| fru-M-000090 | 559 | FruMARCM-M001731_seg001 | fru-Gal4 | M | 2997.73 | 0.53 |
| fru-M-200166 | 1098 | FruMARCM-M001282_seg001 | fru-Gal4 | M | 3226.25 | 0.59 |
| fru-M-300022 | 2404 | FruMARCM-M000127_seg001 | fru-Gal4 | M | 2643.70 | 0.46 |
| 5-HT1B-M-400014 | 2522 | 5HT1bMARCM-M000063_seg001 | 5-HT1B-Gal4 | M | 2908.69 | 0.55 |
| 5-HT1B-M-400016 | 2524 | 5HT1bMARCM-M000065_seg001 | 5-HT1B-Gal4 | M | 3404.95 | 0.64 |
| 5-HT1B-M-200000 | 2538 | 5HT1bMARCM-M000077_seg001 | 5-HT1B-Gal4 | M | 2578.14 | 0.48 |
| fru-F-200163 | 3776 | FruMARCM-F002404_seg002 | fru-Gal4 | F | 2589.91 | 0.48 |
| Cha-F-000240 | 5101 | ChaMARCM-F001160_seg001 | Cha-Gal4 | F | 2260.05 | 0.45 |
| Cha-F-200263 | 5186 | ChaMARCM-F001223_seg001 | Cha-Gal4 | F | 2975.69 | 0.55 |
| Gad1-F-300001 | 6016 | GadMARCM-F002_seg2 | Gad1-Gal4 | F | 3298.82 | 0.60 |
| Cha-F-300125 | 7225 | ChaMARCM-F000390_seg003 | Cha-Gal4 | F | 2927.48 | 0.54 |
| Cha-F-700095 | 7228 | ChaMARCM-F000404_seg001 | Cha-Gal4 | F | 2690.38 | 0.50 |
| Cha-F-400089 | 7249 | ChaMARCM-F000421_seg001 | Cha-Gal4 | F | 3293.31 | 0.56 |
| fru-F-100037 | 7816 | FruMARCM-F000710_seg002 | fru-Gal4 | F | 3139.56 | 0.54 |
| Gad1-F-000045 | 9518 | GadMARCM-F000404_seg001 | Gad1-Gal4 | F | 3159.45 | 0.59 |
| Gad1-F-600076 | 9669 | GadMARCM-F000521_seg002 | Gad1-Gal4 | F | 3115.96 | 0.56 |
| fru-F-400053 | 9773 | FruMARCM-F000282_seg001 | fru-Gal4 | F | 2632.82 | 0.49 |
| fru-F-200046 | 9869 | FruMARCM-F000453_seg001 | fru-Gal4 | F | 3066.66 | 0.51 |
| Gad1-F-200072 | 11290 | GadMARCM-F000262_seg001 | Gad1-Gal4 | F | 3116.14 | 0.54 |
| fru-F-700006 | 11449 | FruMARCM-F000066_seg002 | fru-Gal4 | F | 2897.88 | 0.51 |
| fru-F-200015 | 11507 | FruMARCM-F000170_seg001 | fru-Gal4 | F | 3051.40 | 0.55 |