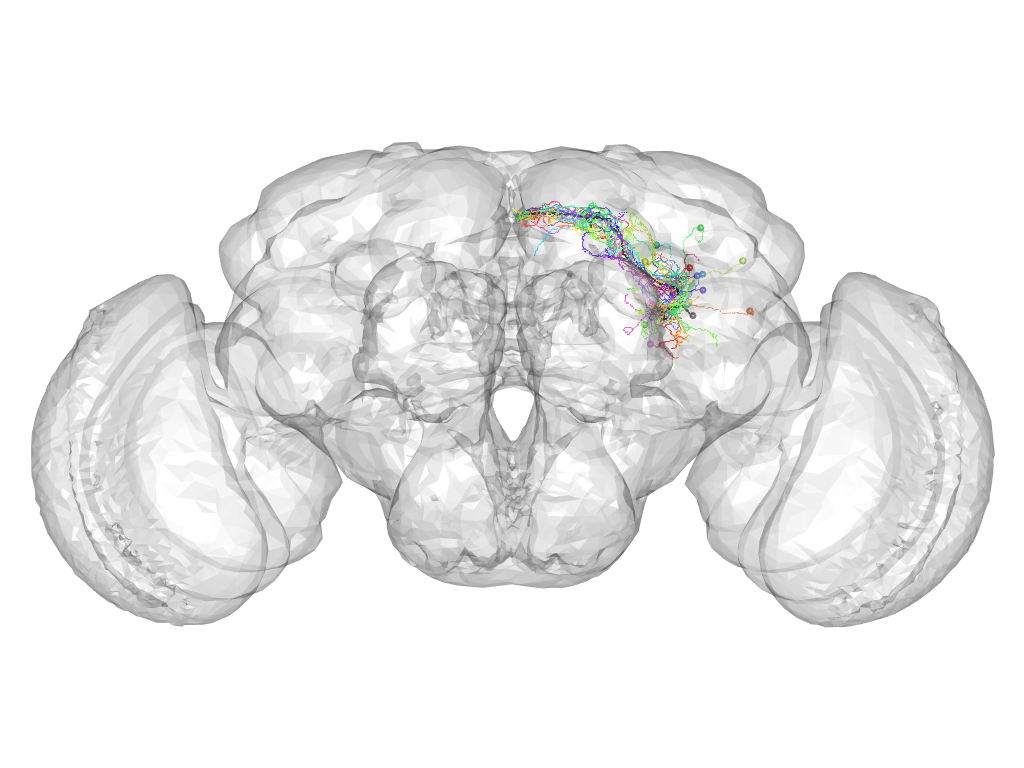
Cluster 504
Part of supercluster XV
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (fru-F-000035), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 404 | fru-F-000048 | XV | 0.62 |
| 526 | Cha-F-400047 | XV | 0.58 |
| 115 | fru-M-000031 | XV | 0.57 |
| 69 | fru-M-000147 | XV | 0.54 |
| 51 | fru-M-000088 | XV | 0.47 |
Cluster composition
This cluster has 18 neurons. The exemplar of this cluster (fru-F-000035) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| fru-F-000035 | 7896 | FruMARCM-F000826_seg001 | fru-Gal4 | F | 4168.46 | 1.00 |
| fru-M-000164 | 33 | FruMARCM-M002230_seg003 | fru-Gal4 | M | 2853.96 | 0.59 |
| fru-M-000093 | 562 | FruMARCM-M001737_seg001 | fru-Gal4 | M | 2712.46 | 0.51 |
| fru-M-000048 | 1130 | FruMARCM-M001325_seg001 | fru-Gal4 | M | 2168.66 | 0.43 |
| fru-M-200009 | 2328 | FruMARCM-M000016_seg001 | fru-Gal4 | M | 2769.71 | 0.62 |
| fru-M-900015 | 2418 | FruMARCM-M000139_seg001 | fru-Gal4 | M | 2715.51 | 0.56 |
| Cha-F-100187 | 4936 | ChaMARCM-F001042_seg001 | Cha-Gal4 | F | 2668.43 | 0.56 |
| Cha-F-000228 | 5034 | ChaMARCM-F001119_seg001 | Cha-Gal4 | F | 2782.09 | 0.58 |
| fru-F-000064 | 6377 | FruMARCM-F001514_seg001 | fru-Gal4 | F | 2558.45 | 0.58 |
| fru-F-400102 | 7868 | FruMARCM-F000793_seg002 | fru-Gal4 | F | 2812.60 | 0.60 |
| Cha-F-200029 | 8113 | ChaMARCM-F000165_seg001 | Cha-Gal4 | F | 2536.53 | 0.62 |
| Gad1-F-600066 | 9652 | GadMARCM-F000504_seg001 | Gad1-Gal4 | F | 2537.63 | 0.53 |
| fru-F-600006 | 9771 | FruMARCM-F000280_seg001 | fru-Gal4 | F | 2710.80 | 0.58 |
| fru-F-000016 | 9800 | FruMARCM-F000348_seg001 | fru-Gal4 | F | 2460.12 | 0.53 |
| Gad1-F-900005 | 10616 | GadMARCM-F000122_seg001 | Gad1-Gal4 | F | 2995.16 | 0.69 |
| fru-F-000007 | 11541 | FruMARCM-F000198_seg001 | fru-Gal4 | F | 2550.26 | 0.56 |
| Gad1-F-300009 | 12615 | GadMARCM-F000019_seg001 | Gad1-Gal4 | F | 2580.57 | 0.57 |
| VGlut-F-000342 | 15019 | DvGlutMARCM-F1359_seg1 | VGlut-Gal4 | F | 2350.16 | 0.55 |