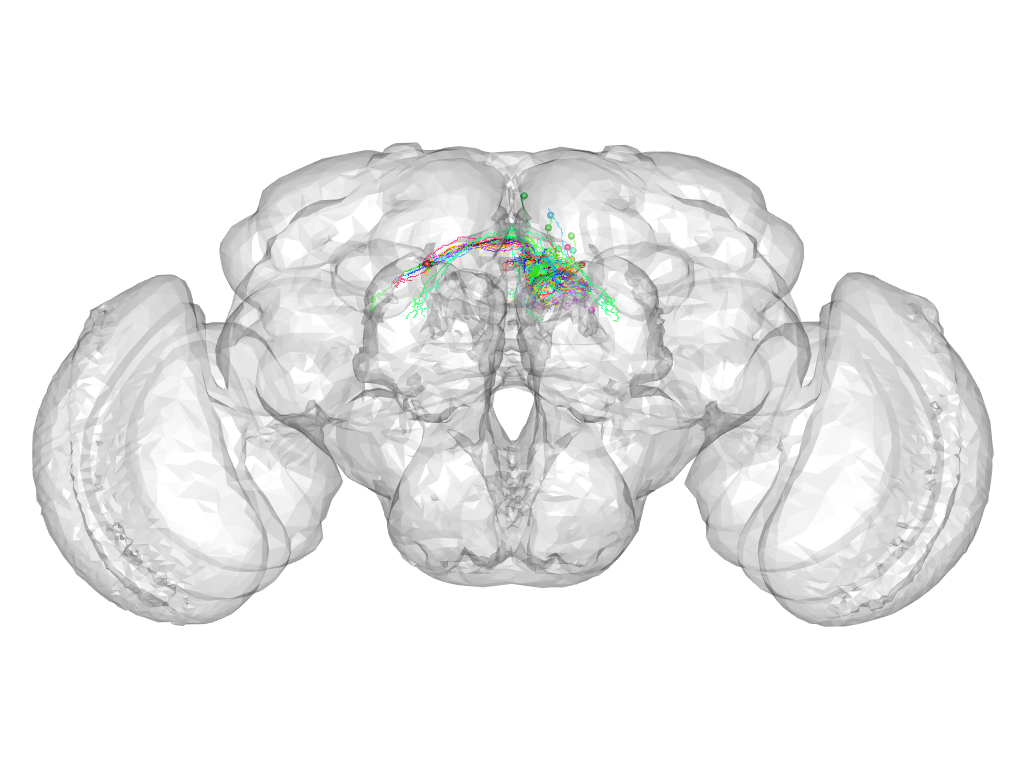
Cluster 405
Part of supercluster XIV
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (fru-F-600072), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 1028 | VGlut-F-000067 | XIV | 0.52 |
| 684 | VGlut-F-200417 | XIV | 0.50 |
| 631 | fru-F-500016 | XIV | 0.47 |
| 964 | VGlut-F-200325 | XIV | 0.46 |
| 77 | fru-M-000157 | XIV | 0.45 |
Cluster composition
This cluster has 25 neurons. The exemplar of this cluster (fru-F-600072) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| fru-F-600072 | 6287 | FruMARCM-F001337_seg001 | fru-Gal4 | F | 7459.95 | 1.00 |
| fru-M-500153 | 469 | FruMARCM-M001574_seg001 | fru-Gal4 | M | 5712.48 | 0.77 |
| fru-M-500168 | 592 | FruMARCM-M001786_seg001 | fru-Gal4 | M | 5662.88 | 0.73 |
| fru-M-300132 | 1045 | FruMARCM-M001226_seg001 | fru-Gal4 | M | 5673.06 | 0.74 |
| fru-M-600049 | 1170 | FruMARCM-M001355_seg001 | fru-Gal4 | M | 5612.45 | 0.74 |
| fru-M-500141 | 1308 | FruMARCM-M001527_seg001 | fru-Gal4 | M | 5570.50 | 0.76 |
| fru-M-700019 | 1766 | FruMARCM-M000286_seg001 | fru-Gal4 | M | 5719.73 | 0.75 |
| fru-F-400225 | 3743 | FruMARCM-F002336_seg001 | fru-Gal4 | F | 5742.59 | 0.75 |
| fru-F-200170 | 3794 | FruMARCM-F002471_seg001 | fru-Gal4 | F | 5387.31 | 0.68 |
| Cha-F-800063 | 3915 | ChaMARCM-F001574_seg001 | Cha-Gal4 | F | 3251.41 | 0.39 |
| Gad1-F-900053 | 4035 | GadMARCM-F000640_seg001 | Gad1-Gal4 | F | 4881.31 | 0.55 |
| Cha-F-000340 | 4354 | ChaMARCM-F001414_seg001 | Cha-Gal4 | F | 4757.61 | 0.58 |
| fru-F-500233 | 4708 | FruMARCM-F002000_seg001 | fru-Gal4 | F | 5631.00 | 0.73 |
| fru-F-500237 | 4716 | FruMARCM-F002042_seg001 | fru-Gal4 | F | 5690.99 | 0.74 |
| fru-F-400135 | 6191 | FruMARCM-F001121_seg001 | fru-Gal4 | F | 5673.94 | 0.71 |
| fru-F-400136 | 6192 | FruMARCM-F001121_seg002 | fru-Gal4 | F | 5358.43 | 0.71 |
| fru-F-500164 | 6212 | FruMARCM-F001136_seg002 | fru-Gal4 | F | 5618.48 | 0.71 |
| fru-F-800045 | 6364 | FruMARCM-F001505_seg001 | fru-Gal4 | F | 5799.75 | 0.75 |
| fru-F-700106 | 7352 | FruMARCM-F000985_seg001 | fru-Gal4 | F | 5248.51 | 0.70 |
| fru-F-500129 | 7805 | FruMARCM-F000703_seg001 | fru-Gal4 | F | 5414.04 | 0.72 |
| Gad1-F-500096 | 9725 | GadMARCM-F000564_seg001 | Gad1-Gal4 | F | 2624.68 | 0.39 |
| fru-F-400068 | 9884 | FruMARCM-F000478_seg001 | fru-Gal4 | F | 5594.13 | 0.75 |
| fru-F-400077 | 9938 | FruMARCM-F000514_seg002 | fru-Gal4 | F | 5637.24 | 0.73 |
| fru-F-500041 | 9955 | FruMARCM-F000547_seg002 | fru-Gal4 | F | 5480.48 | 0.71 |
| Tdc2-F-600000 | 10349 | dTdc2MARCM-F000211_seg001 | Tdc2-Gal4 | F | 5188.59 | 0.67 |