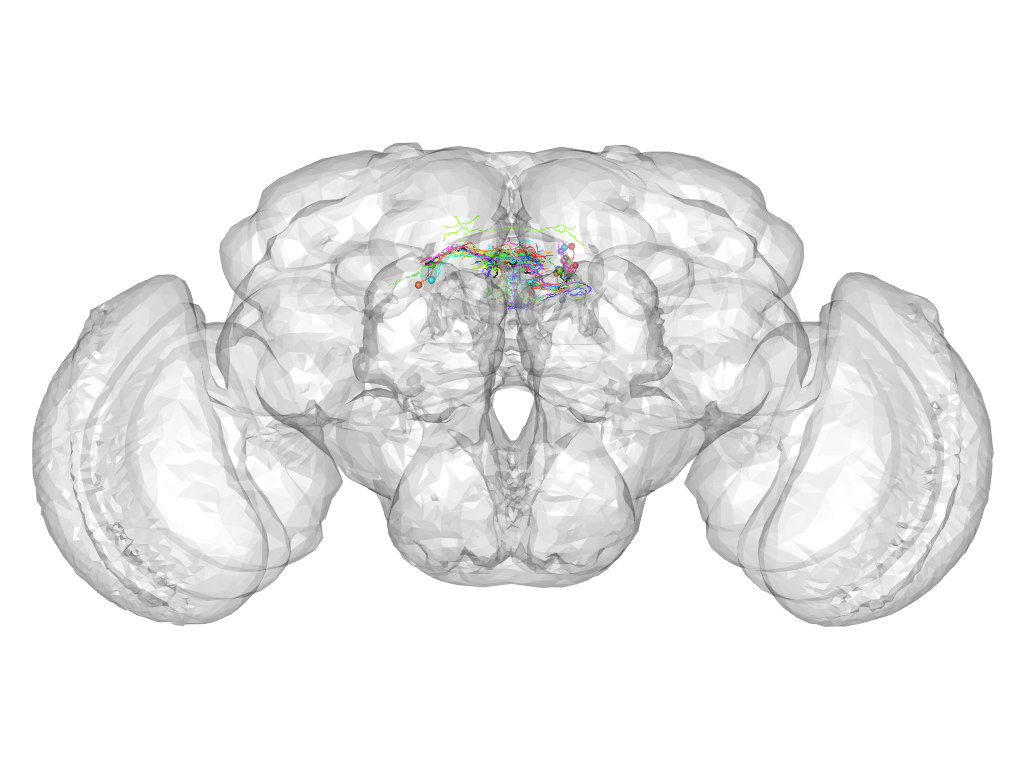
Cluster 77
Part of supercluster XIV
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (fru-M-000157), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 354 | Cha-F-100235 | XIV | 0.67 |
| 126 | fru-M-400032 | XIV | 0.62 |
| 839 | Tdc2-F-300003 | XIV | 0.61 |
| 672 | Gad1-F-300027 | XIV | 0.55 |
| 20 | fru-M-000181 | XIV | 0.50 |
| 1028 | VGlut-F-000067 | XIV | 0.50 |
Cluster composition
This cluster has 21 neurons. The exemplar of this cluster (fru-M-000157) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| fru-M-000157 | 864 | FruMARCM-M002120_seg001 | fru-Gal4 | M | 8393.86 | 1.00 |
| fru-M-200223 | 651 | FruMARCM-M001862_seg001 | fru-Gal4 | M | 6221.14 | 0.72 |
| fru-M-000158 | 868 | FruMARCM-M002123_seg001 | fru-Gal4 | M | 5819.12 | 0.71 |
| fru-M-500067 | 1374 | FruMARCM-M000883_seg001 | fru-Gal4 | M | 6129.91 | 0.73 |
| fru-M-300113 | 1503 | FruMARCM-M001055_seg001 | fru-Gal4 | M | 5864.10 | 0.74 |
| npf-M-700000 | 2021 | npfMARCM-M000013_seg001 | npf-GAL4 | M | 4085.32 | 0.51 |
| Gad1-F-100076 | 4057 | GadMARCM-F000655_seg001 | Gad1-Gal4 | F | 5548.39 | 0.59 |
| fru-F-500207 | 4550 | FruMARCM-F001660_seg001 | fru-Gal4 | F | 6360.72 | 0.75 |
| fru-F-500210 | 4585 | FruMARCM-F001700_seg001 | fru-Gal4 | F | 6258.28 | 0.75 |
| fru-F-700173 | 4818 | FruMARCM-F002185_seg001 | fru-Gal4 | F | 6067.07 | 0.73 |
| fru-F-700124 | 6239 | FruMARCM-F001218_seg001 | fru-Gal4 | F | 5512.44 | 0.68 |
| fru-F-400110 | 7314 | FruMARCM-F000953_seg001 | fru-Gal4 | F | 6003.44 | 0.71 |
| fru-F-400095 | 7859 | FruMARCM-F000767_seg001 | fru-Gal4 | F | 6016.62 | 0.74 |
| fru-F-400101 | 7867 | FruMARCM-F000793_seg001 | fru-Gal4 | F | 5937.42 | 0.72 |
| npf-F-400005 | 9459 | npfMARCM-F000046_seg001 | npf-GAL4 | F | 4798.43 | 0.60 |
| Gad1-F-100035 | 9525 | GadMARCM-F000408_seg001 | Gad1-Gal4 | F | 4313.17 | 0.55 |
| Gad1-F-500076 | 9530 | GadMARCM-F000413_seg001 | Gad1-Gal4 | F | 3788.62 | 0.53 |
| Gad1-F-400083 | 9562 | GadMARCM-F000437_seg001 | Gad1-Gal4 | F | 5317.80 | 0.71 |
| Gad1-F-400106 | 9717 | GadMARCM-F000559_seg001 | Gad1-Gal4 | F | 5177.09 | 0.70 |
| Gad1-F-600004 | 10578 | GadMARCM-F000098_seg001 | Gad1-Gal4 | F | 6090.83 | 0.74 |
| Trh-F-500007 | 13854 | TPHMARCM-125F_seg2 | Trh-Gal4 | F | 5358.99 | 0.69 |