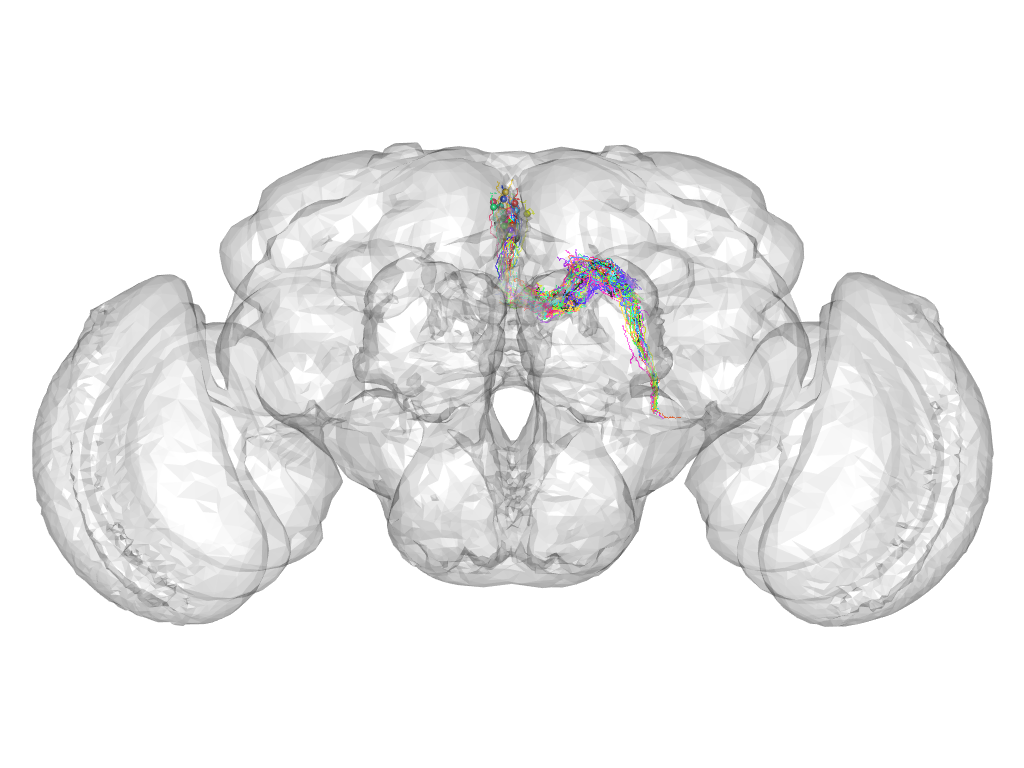
Cluster 346
Part of supercluster XIV
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (Cha-F-100230), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 726 | Gad1-F-300066 | XIV | 0.54 |
| 667 | TH-F-300071 | XIV | 0.50 |
| 191 | TH-M-200018 | XIV | 0.43 |
| 1028 | VGlut-F-000067 | XIV | 0.42 |
| 204 | Trh-M-100025 | XIV | 0.42 |
Cluster composition
This cluster has 23 neurons. The exemplar of this cluster (Cha-F-100230) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| Cha-F-100230 | 5165 | ChaMARCM-F001207_seg001 | Cha-Gal4 | F | 19407.25 | 1.00 |
| Cha-F-500186 | 3846 | ChaMARCM-F001519_seg001 | Cha-Gal4 | F | 14332.78 | 0.73 |
| Gad1-F-100083 | 4122 | GadMARCM-F000708_seg001 | Gad1-Gal4 | F | 15648.36 | 0.77 |
| Cha-F-000293 | 4183 | ChaMARCM-F001288_seg001 | Cha-Gal4 | F | 14771.36 | 0.76 |
| Cha-F-100268 | 4227 | ChaMARCM-F001316_seg001 | Cha-Gal4 | F | 15591.85 | 0.78 |
| Cha-F-200191 | 4858 | ChaMARCM-F000980_seg001 | Cha-Gal4 | F | 15285.03 | 0.78 |
| Cha-F-300179 | 5257 | ChaMARCM-F000656_seg001 | Cha-Gal4 | F | 14719.60 | 0.77 |
| Cha-F-900030 | 5404 | ChaMARCM-F000758_seg001 | Cha-Gal4 | F | 14252.60 | 0.73 |
| Cha-F-600104 | 5420 | ChaMARCM-F000772_seg001 | Cha-Gal4 | F | 14746.61 | 0.77 |
| Cha-F-800050 | 5677 | ChaMARCM-F000939_seg001 | Cha-Gal4 | F | 15544.47 | 0.80 |
| Cha-F-400135 | 7151 | ChaMARCM-F000613_seg001 | Cha-Gal4 | F | 14645.16 | 0.75 |
| Cha-F-700106 | 7578 | ChaMARCM-F000437_seg001 | Cha-Gal4 | F | 14534.27 | 0.75 |
| Cha-F-200076 | 7596 | ChaMARCM-F000448_seg001 | Cha-Gal4 | F | 13697.19 | 0.72 |
| Cha-F-200084 | 7636 | ChaMARCM-F000472_seg001 | Cha-Gal4 | F | 13038.58 | 0.69 |
| Cha-F-300043 | 8138 | ChaMARCM-F000177_seg001 | Cha-Gal4 | F | 14296.26 | 0.73 |
| Cha-F-000014 | 10154 | ChaMARCM-F000089_seg001 | Cha-Gal4 | F | 14081.38 | 0.74 |
| Tdc2-F-300033 | 10334 | dTdc2MARCM-F000196_seg001 | Tdc2-Gal4 | F | 10481.03 | 0.57 |
| Tdc2-F-300034 | 10335 | dTdc2MARCM-F000197_seg001 | Tdc2-Gal4 | F | 9195.07 | 0.53 |
| Tdc2-F-300038 | 10339 | dTdc2MARCM-F000200_seg001 | Tdc2-Gal4 | F | 10421.99 | 0.53 |
| Gad1-F-700012 | 10635 | GadMARCM-F000135_seg002 | Gad1-Gal4 | F | 13966.10 | 0.73 |
| Gad1-F-500042 | 10659 | GadMARCM-F000172_seg001 | Gad1-Gal4 | F | 14260.33 | 0.71 |
| Gad1-F-100009 | 10722 | GadMARCM-F000049_seg002 | Gad1-Gal4 | F | 13818.95 | 0.70 |
| Gad1-F-200093 | 11339 | GadMARCM-F000306_seg001 | Gad1-Gal4 | F | 14771.19 | 0.78 |