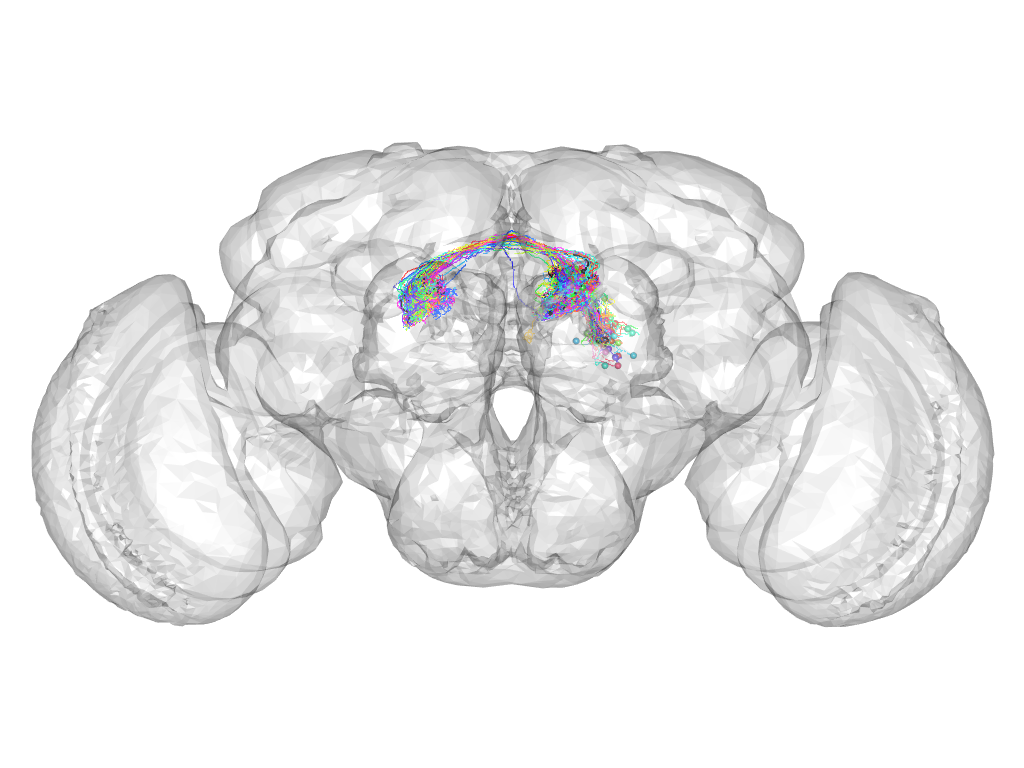
Cluster 265
Part of supercluster XIV
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (Cha-F-100295), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 245 | Gad1-F-200134 | XIV | 0.69 |
| 1028 | VGlut-F-000067 | XIV | 0.56 |
| 964 | VGlut-F-200325 | XIV | 0.54 |
| 32 | fru-M-500240 | XIV | 0.53 |
| 414 | fru-F-400152 | XIV | 0.52 |
| 325 | Cha-F-100151 | XIV | 0.51 |
| 840 | Tdc2-F-100009 | XIV | 0.51 |
Cluster composition
This cluster has 43 neurons. The exemplar of this cluster (Cha-F-100295) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| Cha-F-100295 | 4278 | ChaMARCM-F001354_seg001 | Cha-Gal4 | F | 26069.95 | 1.00 |
| Cha-F-500188 | 3848 | ChaMARCM-F001520_seg001 | Cha-Gal4 | F | 20126.81 | 0.78 |
| Gad1-F-100065 | 3947 | GadMARCM-F000577_seg001 | Gad1-Gal4 | F | 20789.22 | 0.79 |
| Gad1-F-000059 | 3987 | GadMARCM-F000608_seg001 | Gad1-Gal4 | F | 19744.17 | 0.77 |
| Gad1-F-200145 | 4097 | GadMARCM-F000687_seg001 | Gad1-Gal4 | F | 20218.31 | 0.79 |
| Gad1-F-000091 | 4103 | GadMARCM-F000691_seg001 | Gad1-Gal4 | F | 19760.74 | 0.76 |
| Gad1-F-000100 | 4112 | GadMARCM-F000697_seg001 | Gad1-Gal4 | F | 20211.12 | 0.77 |
| Cha-F-200277 | 4143 | ChaMARCM-F001260_seg001 | Cha-Gal4 | F | 20557.20 | 0.80 |
| Cha-F-200364 | 4389 | ChaMARCM-F001440_seg002 | Cha-Gal4 | F | 18339.26 | 0.72 |
| Cha-F-300309 | 4438 | ChaMARCM-F001481_seg001 | Cha-Gal4 | F | 20310.36 | 0.78 |
| Cha-F-000381 | 4468 | ChaMARCM-F001508_seg001 | Cha-Gal4 | F | 19688.56 | 0.76 |
| Cha-F-200198 | 4865 | ChaMARCM-F000986_seg002 | Cha-Gal4 | F | 19640.21 | 0.77 |
| Cha-F-100130 | 4875 | ChaMARCM-F000995_seg001 | Cha-Gal4 | F | 19696.74 | 0.77 |
| Cha-F-000195 | 4983 | ChaMARCM-F001078_seg001 | Cha-Gal4 | F | 20717.94 | 0.80 |
| Cha-F-000203 | 4996 | ChaMARCM-F001090_seg001 | Cha-Gal4 | F | 20214.93 | 0.77 |
| Cha-F-000254 | 5172 | ChaMARCM-F001213_seg001 | Cha-Gal4 | F | 19641.78 | 0.77 |
| Cha-F-100242 | 5217 | ChaMARCM-F001250_seg001 | Cha-Gal4 | F | 20511.61 | 0.79 |
| Cha-F-000134 | 5362 | ChaMARCM-F000731_seg001 | Cha-Gal4 | F | 19007.43 | 0.75 |
| Cha-F-300234 | 5504 | ChaMARCM-F000823_seg001 | Cha-Gal4 | F | 16448.56 | 0.66 |
| Cha-F-700150 | 5519 | ChaMARCM-F000832_seg001 | Cha-Gal4 | F | 20083.69 | 0.78 |
| Cha-F-000147 | 5536 | ChaMARCM-F000842_seg001 | Cha-Gal4 | F | 20095.68 | 0.77 |
| Cha-F-000155 | 5583 | ChaMARCM-F000877_seg001 | Cha-Gal4 | F | 19963.27 | 0.77 |
| Cha-F-300253 | 5667 | ChaMARCM-F000935_seg001 | Cha-Gal4 | F | 20313.26 | 0.78 |
| Cha-F-100065 | 7092 | ChaMARCM-F000570_seg001 | Cha-Gal4 | F | 19437.87 | 0.76 |
| Cha-F-700138 | 7115 | ChaMARCM-F000588_seg001 | Cha-Gal4 | F | 16709.99 | 0.69 |
| Cha-F-000103 | 7161 | ChaMARCM-F000618_seg001 | Cha-Gal4 | F | 19505.16 | 0.76 |
| Cha-F-100033 | 7545 | ChaMARCM-F000398_seg001 | Cha-Gal4 | F | 20502.02 | 0.80 |
| Cha-F-500108 | 7627 | ChaMARCM-F000466_seg002 | Cha-Gal4 | F | 18228.53 | 0.70 |
| Cha-F-300132 | 7631 | ChaMARCM-F000469_seg001 | Cha-Gal4 | F | 20245.03 | 0.77 |
| Cha-F-700126 | 7654 | ChaMARCM-F000485_seg001 | Cha-Gal4 | F | 19167.01 | 0.74 |
| Cha-F-600083 | 7732 | ChaMARCM-F000541_seg001 | Cha-Gal4 | F | 15635.18 | 0.65 |
| Cha-F-300100 | 8352 | ChaMARCM-F000301_seg001 | Cha-Gal4 | F | 15942.88 | 0.67 |
| Gad1-F-800025 | 9493 | GadMARCM-F000386_seg001 | Gad1-Gal4 | F | 19623.69 | 0.76 |
| Gad1-F-300123 | 9618 | GadMARCM-F000479_seg001 | Gad1-Gal4 | F | 18707.65 | 0.74 |
| Gad1-F-300132 | 9711 | GadMARCM-F000553_seg001 | Gad1-Gal4 | F | 18927.51 | 0.75 |
| Gad1-F-500093 | 9720 | GadMARCM-F000562_seg001 | Gad1-Gal4 | F | 19276.94 | 0.76 |
| Cha-F-400017 | 10118 | ChaMARCM-F000070_seg001 | Cha-Gal4 | F | 19282.18 | 0.75 |
| Cha-F-200023 | 10161 | ChaMARCM-F000093_seg002 | Cha-Gal4 | F | 19096.87 | 0.75 |
| Gad1-F-500035 | 10519 | GadMARCM-F000161_seg001 | Gad1-Gal4 | F | 19893.28 | 0.77 |
| Gad1-F-200044 | 10685 | GadMARCM-F000187_seg001 | Gad1-Gal4 | F | 20402.47 | 0.79 |
| Gad1-F-800013 | 10688 | GadMARCM-F000189_seg001 | Gad1-Gal4 | F | 20507.77 | 0.79 |
| Gad1-F-300063 | 11226 | GadMARCM-F000209_seg001 | Gad1-Gal4 | F | 20453.38 | 0.78 |
| Gad1-F-400032 | 11269 | GadMARCM-F000246_seg002 | Gad1-Gal4 | F | 20143.50 | 0.78 |