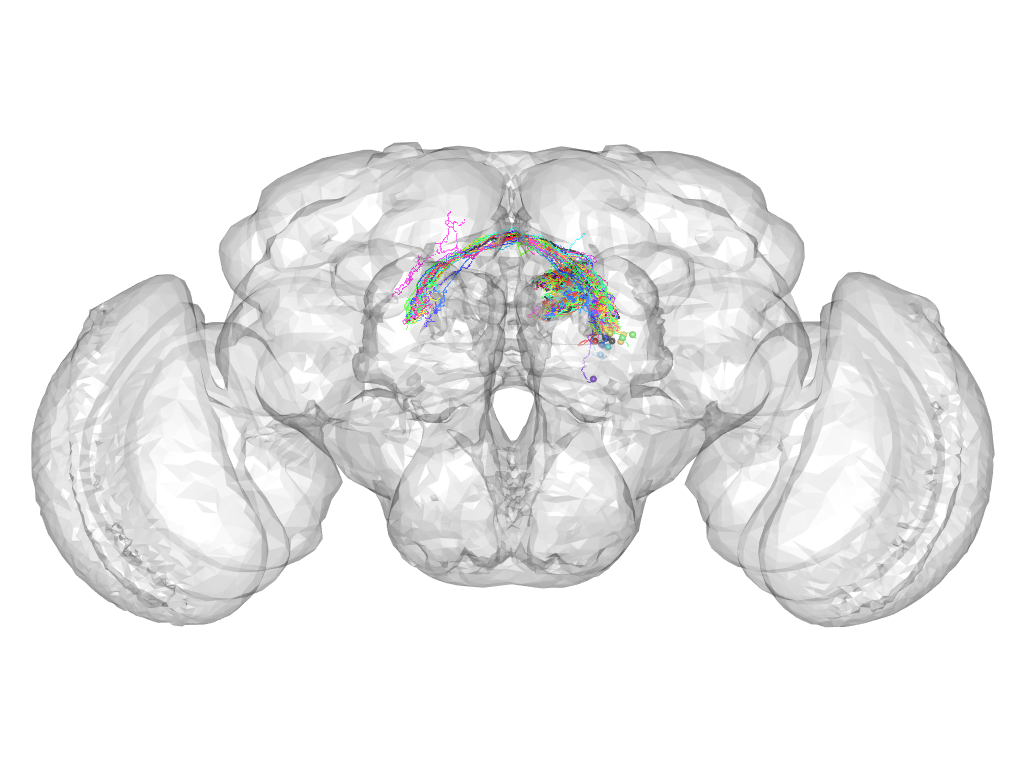
Cluster 245
Part of supercluster XIV
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (Gad1-F-200134), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 265 | Cha-F-100295 | XIV | 0.51 |
| 964 | VGlut-F-200325 | XIV | 0.41 |
| 325 | Cha-F-100151 | XIV | 0.40 |
| 1028 | VGlut-F-000067 | XIV | 0.39 |
| 414 | fru-F-400152 | XIV | 0.36 |
Cluster composition
This cluster has 27 neurons. The exemplar of this cluster (Gad1-F-200134) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| Gad1-F-200134 | 3989 | GadMARCM-F000609_seg001 | Gad1-Gal4 | F | 22892.35 | 1.00 |
| Cha-F-300332 | 3887 | ChaMARCM-F001549_seg002 | Cha-Gal4 | F | 17571.95 | 0.77 |
| Cha-F-200283 | 4192 | ChaMARCM-F001294_seg001 | Cha-Gal4 | F | 17911.05 | 0.79 |
| Cha-F-000354 | 4410 | ChaMARCM-F001459_seg001 | Cha-Gal4 | F | 17792.47 | 0.77 |
| Cha-F-000356 | 4412 | ChaMARCM-F001461_seg001 | Cha-Gal4 | F | 17726.83 | 0.77 |
| Cha-F-100119 | 4851 | ChaMARCM-F000975_seg001 | Cha-Gal4 | F | 18043.26 | 0.79 |
| Cha-F-200202 | 4916 | ChaMARCM-F001027_seg001 | Cha-Gal4 | F | 17715.50 | 0.78 |
| Cha-F-200205 | 4960 | ChaMARCM-F001058_seg001 | Cha-Gal4 | F | 17989.17 | 0.79 |
| Cha-F-000190 | 4969 | ChaMARCM-F001066_seg001 | Cha-Gal4 | F | 17861.10 | 0.76 |
| Cha-F-000211 | 5016 | ChaMARCM-F001106_seg001 | Cha-Gal4 | F | 17478.57 | 0.77 |
| Cha-F-100225 | 5157 | ChaMARCM-F001202_seg001 | Cha-Gal4 | F | 17977.17 | 0.78 |
| Cha-F-500140 | 5293 | ChaMARCM-F000681_seg001 | Cha-Gal4 | F | 16902.89 | 0.74 |
| Cha-F-300215 | 5450 | ChaMARCM-F000788_seg001 | Cha-Gal4 | F | 17249.94 | 0.75 |
| Cha-F-300223 | 5484 | ChaMARCM-F000809_seg001 | Cha-Gal4 | F | 17459.37 | 0.77 |
| Gad1-F-300085 | 6149 | GadMARCM-F000338_seg001 | Gad1-Gal4 | F | 17842.15 | 0.78 |
| Cha-F-700135 | 7101 | ChaMARCM-F000577_seg001 | Cha-Gal4 | F | 17066.41 | 0.75 |
| Cha-F-700089 | 7530 | ChaMARCM-F000378_seg002 | Cha-Gal4 | F | 15714.12 | 0.71 |
| Cha-F-100047 | 7667 | ChaMARCM-F000496_seg001 | Cha-Gal4 | F | 14169.32 | 0.65 |
| Cha-F-700130 | 7743 | ChaMARCM-F000548_seg001 | Cha-Gal4 | F | 18059.22 | 0.79 |
| Cha-F-200051 | 8241 | ChaMARCM-F000237_seg001 | Cha-Gal4 | F | 15645.44 | 0.68 |
| Cha-F-200068 | 8283 | ChaMARCM-F000260_seg001 | Cha-Gal4 | F | 17627.94 | 0.76 |
| Gad1-F-600081 | 9699 | GadMARCM-F000544_seg001 | Gad1-Gal4 | F | 16574.93 | 0.74 |
| Cha-F-400012 | 10081 | ChaMARCM-F000041_seg001 | Cha-Gal4 | F | 17422.68 | 0.76 |
| Cha-F-300014 | 10100 | ChaMARCM-F000055_seg002 | Cha-Gal4 | F | 12929.42 | 0.56 |
| Gad1-F-200060 | 11238 | GadMARCM-F000220_seg001 | Gad1-Gal4 | F | 17069.23 | 0.75 |
| Cha-F-600001 | 11595 | ChaMARCM-F000026_seg001 | Cha-Gal4 | F | 17765.51 | 0.78 |
| Gad1-F-200001 | 13179 | GadMARCM-F014_seg1 | Gad1-Gal4 | F | 17278.63 | 0.75 |