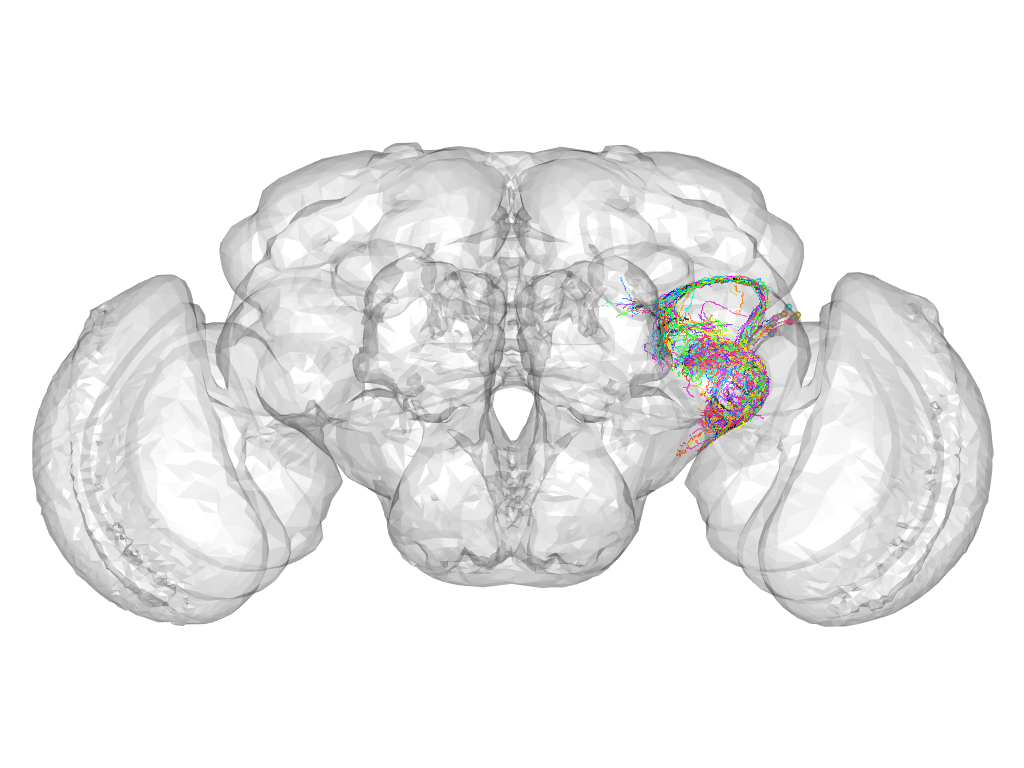
Cluster 212
Part of supercluster X
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (Trh-M-700003), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 805 | Trh-F-500193 | X | 0.65 |
| 876 | Trh-F-100023 | X | 0.63 |
| 200 | TH-M-000060 | X | 0.60 |
| 865 | TH-F-000025 | X | 0.57 |
| 268 | Cha-F-200340 | X | 0.48 |
Cluster composition
This cluster has 25 neurons. The exemplar of this cluster (Trh-M-700003) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| Trh-M-700003 | 3654 | TPHMARCM-669M_seg1 | Trh-Gal4 | M | 12015.64 | 1.00 |
| Trh-M-500126 | 930 | TPHMARCM-M001499_seg001 | Trh-Gal4 | M | 8404.52 | 0.70 |
| Trh-M-500181 | 2086 | TPHMARCM-M001587_seg001 | Trh-Gal4 | M | 8413.24 | 0.66 |
| Trh-M-500155 | 2547 | TPHMARCM-M001526_seg001 | Trh-Gal4 | M | 8510.57 | 0.68 |
| Trh-M-500145 | 2742 | TPHMARCM-M001508_seg001 | Trh-Gal4 | M | 8887.24 | 0.70 |
| Trh-M-500150 | 2755 | TPHMARCM-M001522_seg001 | Trh-Gal4 | M | 8142.66 | 0.69 |
| Trh-M-700040 | 2788 | TPHMARCM-1013M_seg1 | Trh-Gal4 | M | 8104.77 | 0.66 |
| Trh-M-400053 | 2806 | TPHMARCM-1035M_seg1 | Trh-Gal4 | M | 7379.65 | 0.67 |
| Trh-M-400100 | 2878 | TPHMARCM-1123M_seg1 | Trh-Gal4 | M | 8687.76 | 0.69 |
| Trh-F-500070 | 10449 | TPHMARCM-675F_seg1 | Trh-Gal4 | F | 8191.86 | 0.67 |
| Trh-F-700057 | 12412 | TPHMARCM-1205F_seg1 | Trh-Gal4 | F | 8400.15 | 0.69 |
| Trh-F-500119 | 12437 | TPHMARCM-1231F_seg1 | Trh-Gal4 | F | 8187.89 | 0.70 |
| Trh-F-500145 | 12480 | TPHMARCM-1264F_seg1 | Trh-Gal4 | F | 8141.52 | 0.68 |
| Trh-F-500146 | 12481 | TPHMARCM-1265F_seg1 | Trh-Gal4 | F | 8649.07 | 0.71 |
| Trh-F-500214 | 12585 | TPHMARCM-F001379_seg001 | Trh-Gal4 | F | 8526.67 | 0.72 |
| Trh-F-500085 | 13254 | TPHMARCM-741F_seg1 | Trh-Gal4 | F | 8795.30 | 0.70 |
| Trh-F-600040 | 13284 | TPHMARCM-778F_seg1 | Trh-Gal4 | F | 8779.72 | 0.72 |
| Trh-F-600041 | 13288 | TPHMARCM-779F_seg1 | Trh-Gal4 | F | 8204.94 | 0.71 |
| Trh-F-400001 | 13827 | TPHMARCM-048F_seg1 | Trh-Gal4 | F | 8173.98 | 0.66 |
| Trh-F-400004 | 13830 | TPHMARCM-051F_seg1 | Trh-Gal4 | F | 8061.70 | 0.65 |
| Trh-F-400029 | 13948 | TPHMARCM-240F_seg1 | Trh-Gal4 | F | 8738.99 | 0.71 |
| Trh-F-400037 | 13985 | TPHMARCM-275F_seg1 | Trh-Gal4 | F | 8233.84 | 0.70 |
| Trh-F-400041 | 14026 | TPHMARCM-312F_seg1 | Trh-Gal4 | F | 8387.44 | 0.66 |
| Trh-F-400052 | 14140 | TPHMARCM-552F_seg1 | Trh-Gal4 | F | 8868.93 | 0.71 |
| Trh-F-700037 | 14273 | TPHMARCM-820F_seg2 | Trh-Gal4 | F | 8249.54 | 0.66 |