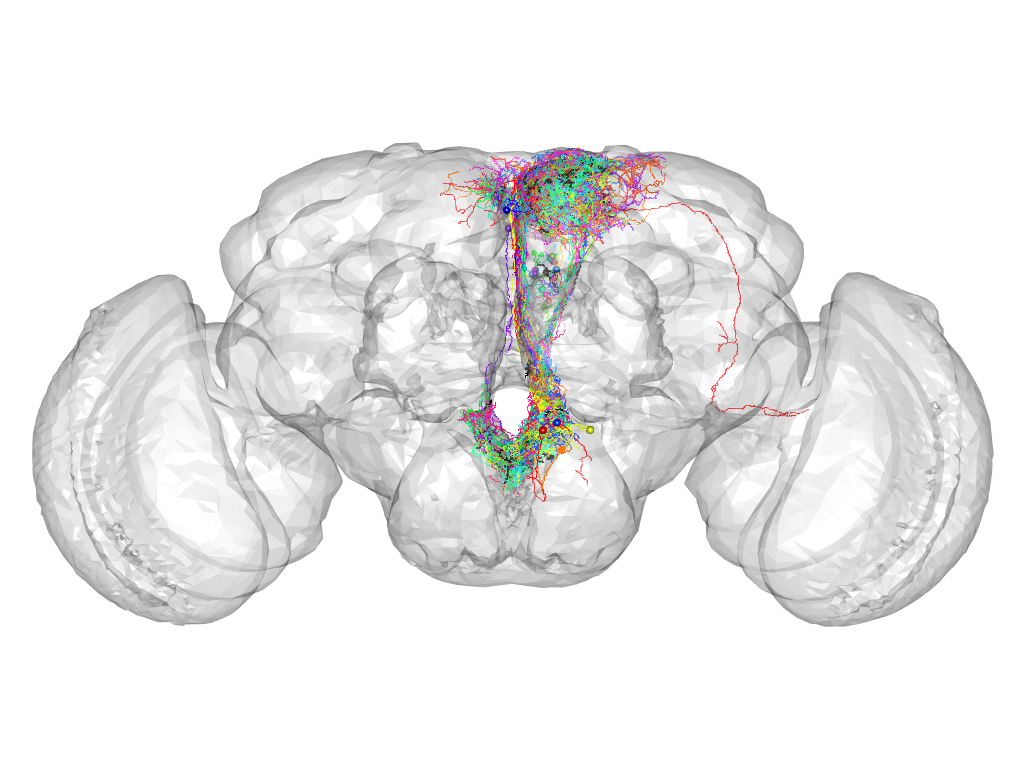
Cluster 206
Part of supercluster XIX
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (Trh-M-000018), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 199 | TH-M-000044 | XIX | 0.61 |
| 95 | fru-M-000064 | XIX | 0.42 |
| 283 | Cha-F-100343 | IX | 0.39 |
| 176 | Trh-M-300108 | IX | 0.31 |
| 698 | VGlut-F-600430 | IX | 0.31 |
Cluster composition
This cluster has 28 neurons. The exemplar of this cluster (Trh-M-000018) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| Trh-M-000018 | 3481 | TPHMARCM-352M_seg2 | Trh-Gal4 | M | 59041.77 | 1.00 |
| fru-M-300250 | 54 | FruMARCM-M002250_seg001 | fru-Gal4 | M | 34955.01 | 0.48 |
| fru-M-500230 | 153 | FruMARCM-M002377_seg003 | fru-Gal4 | M | 30815.23 | 0.52 |
| fru-M-600083 | 631 | FruMARCM-M001842_seg001 | fru-Gal4 | M | 33788.24 | 0.55 |
| fru-M-900029 | 1139 | FruMARCM-M001332_seg001 | fru-Gal4 | M | 27176.29 | 0.58 |
| fru-M-600057 | 1197 | FruMARCM-M001377_seg001 | fru-Gal4 | M | 35591.50 | 0.67 |
| fru-M-200202 | 1294 | FruMARCM-M001487_seg001 | fru-Gal4 | M | 34769.12 | 0.59 |
| fru-M-400095 | 1472 | FruMARCM-M001031_seg001 | fru-Gal4 | M | 26514.46 | 0.57 |
| fru-M-100069 | 1651 | FruMARCM-M000790_seg001 | fru-Gal4 | M | 32897.29 | 0.64 |
| Trh-M-100035 | 1976 | TPHMARCM-0345M_seg001 | Trh-Gal4 | M | 42995.48 | 0.72 |
| Trh-M-200118 | 2034 | TPHMARCM-M001755_seg001 | Trh-Gal4 | M | 43819.95 | 0.73 |
| Trh-M-000060 | 2111 | TPHMARCM-M001609_seg001 | Trh-Gal4 | M | 44074.37 | 0.73 |
| Trh-M-000091 | 2265 | TPHMARCM-M001752_seg001 | Trh-Gal4 | M | 44486.06 | 0.74 |
| Trh-M-000118 | 2304 | TPHMARCM-M001786_seg001 | Trh-Gal4 | M | 43565.47 | 0.73 |
| Trh-M-100005 | 3388 | TPHMARCM-067M_seg1 | Trh-Gal4 | M | 43524.35 | 0.72 |
| Trh-M-100029 | 3469 | TPHMARCM-339M_seg1 | Trh-Gal4 | M | 42623.77 | 0.72 |
| Trh-M-200022 | 3484 | TPHMARCM-357M_seg1 | Trh-Gal4 | M | 43110.71 | 0.73 |
| Cha-F-400243 | 3892 | ChaMARCM-F001554_seg001 | Cha-Gal4 | F | 25618.02 | 0.50 |
| fru-F-200134 | 4636 | FruMARCM-F001774_seg001 | fru-Gal4 | F | 36506.47 | 0.63 |
| Cha-F-000208 | 5013 | ChaMARCM-F001104_seg001 | Cha-Gal4 | F | 23847.58 | 0.54 |
| Cha-F-300227 | 5496 | ChaMARCM-F000817_seg002 | Cha-Gal4 | F | 31942.32 | 0.64 |
| fru-F-400159 | 6367 | FruMARCM-F001507_seg001 | fru-Gal4 | F | 16868.90 | 0.44 |
| VGlut-F-500823 | 6611 | DvGlutMARCM-F004365_seg003 | VGlut-Gal4 | F | 30340.76 | 0.54 |
| VGlut-F-600738 | 6879 | DvGlutMARCM-F004038_seg004 | VGlut-Gal4 | F | 32426.94 | 0.63 |
| fru-F-200054 | 7755 | FruMARCM-F000648_seg001 | fru-Gal4 | F | 29248.56 | 0.52 |
| Trh-F-100024 | 13818 | TPHMARCM-037F_seg1 | Trh-Gal4 | F | 44666.43 | 0.73 |
| Trh-F-200021 | 13944 | TPHMARCM-236F-repeated0811-07_seg1 | Trh-Gal4 | F | 41919.04 | 0.70 |
| Trh-F-100043 | 14041 | TPHMARCM-370F_seg1 | Trh-Gal4 | F | 43503.08 | 0.72 |