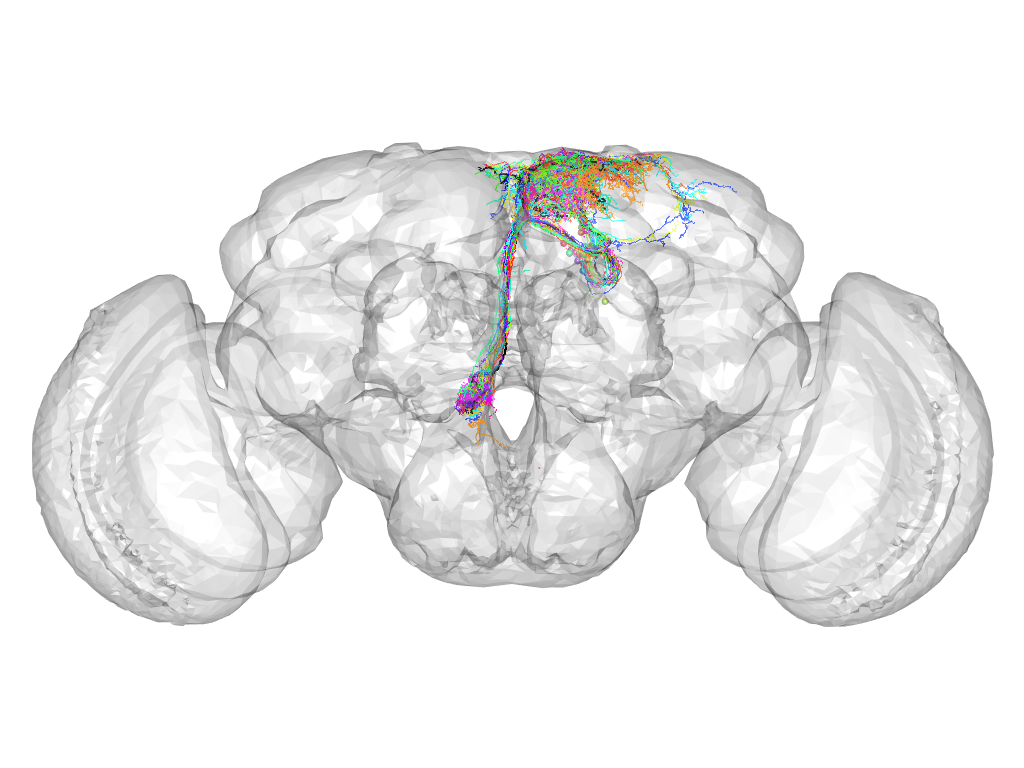
Cluster 95
Part of supercluster XIX
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (fru-M-000064), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 199 | TH-M-000044 | XIX | 0.66 |
| 206 | Trh-M-000018 | XIX | 0.57 |
| 881 | Trh-F-200023 | XIX | 0.51 |
| 117 | fru-M-500059 | XVIII | 0.47 |
| 40 | fru-M-000123 | XIX | 0.45 |
Cluster composition
This cluster has 23 neurons. The exemplar of this cluster (fru-M-000064) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| fru-M-000064 | 1187 | FruMARCM-M001369_seg001 | fru-Gal4 | M | 23359.31 | 1.00 |
| fru-F-800062 | 3722 | FruMARCM-F002220_seg002 | fru-Gal4 | F | 16711.38 | 0.71 |
| fru-F-400227 | 3745 | FruMARCM-F002337_seg001 | fru-Gal4 | F | 17108.09 | 0.70 |
| Cha-F-000300 | 4214 | ChaMARCM-F001303_seg001 | Cha-Gal4 | F | 13099.39 | 0.52 |
| fru-F-400171 | 4536 | FruMARCM-F001644_seg002 | fru-Gal4 | F | 16201.01 | 0.71 |
| fru-F-500209 | 4552 | FruMARCM-F001660_seg003 | fru-Gal4 | F | 14039.01 | 0.53 |
| fru-F-500234 | 4709 | FruMARCM-F002001_seg001 | fru-Gal4 | F | 16599.56 | 0.71 |
| fru-F-500238 | 4717 | FruMARCM-F002042_seg002 | fru-Gal4 | F | 16902.89 | 0.72 |
| fru-F-500249 | 4778 | FruMARCM-F002095_seg001 | fru-Gal4 | F | 16408.59 | 0.73 |
| fru-F-500253 | 4782 | FruMARCM-F002104_seg002 | fru-Gal4 | F | 16120.69 | 0.71 |
| fru-F-100070 | 4806 | FruMARCM-F002149_seg001 | fru-Gal4 | F | 16124.29 | 0.70 |
| fru-F-700174 | 4819 | FruMARCM-F002185_seg002 | fru-Gal4 | F | 17044.99 | 0.73 |
| Cha-F-400197 | 5581 | ChaMARCM-F000874_seg002 | Cha-Gal4 | F | 16553.66 | 0.64 |
| fru-F-600079 | 6308 | FruMARCM-F001427_seg001 | fru-Gal4 | F | 15168.53 | 0.69 |
| fru-F-500180 | 6318 | FruMARCM-F001441_seg003 | fru-Gal4 | F | 14543.78 | 0.68 |
| fru-F-400155 | 6360 | FruMARCM-F001502_seg004 | fru-Gal4 | F | 13445.09 | 0.48 |
| fru-F-700087 | 7299 | FruMARCM-F000928_seg003 | fru-Gal4 | F | 10850.82 | 0.57 |
| fru-F-400049 | 9765 | FruMARCM-F000275_seg002 | fru-Gal4 | F | 13078.52 | 0.62 |
| fru-F-700040 | 9900 | FruMARCM-F000490_seg001 | fru-Gal4 | F | 16284.84 | 0.72 |
| fru-F-600039 | 9993 | FruMARCM-F000580_seg001 | fru-Gal4 | F | 16141.09 | 0.68 |
| fru-F-500068 | 9994 | FruMARCM-F000581_seg001 | fru-Gal4 | F | 16276.81 | 0.69 |
| fru-F-500073 | 9999 | FruMARCM-F000583_seg004 | fru-Gal4 | F | 15363.78 | 0.69 |
| fru-F-500075 | 10001 | FruMARCM-F000584_seg002 | fru-Gal4 | F | 16035.31 | 0.70 |