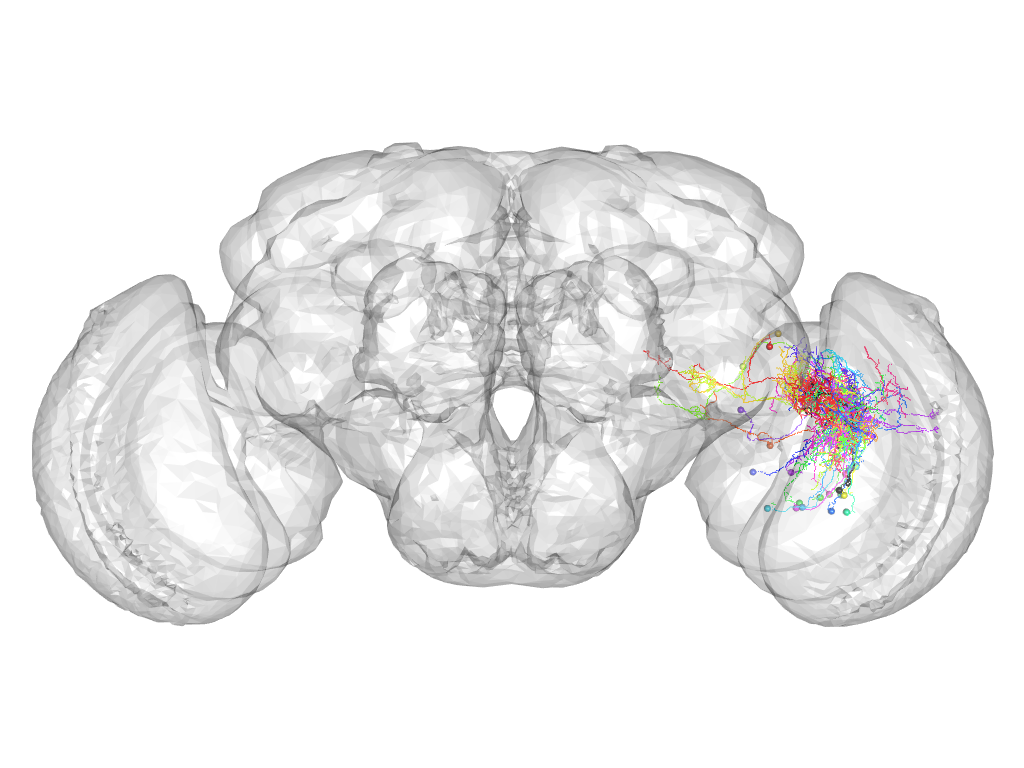
Cluster 1030
Part of supercluster I
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (VGlut-F-000074), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 127 | fru-M-700026 | V | 0.70 |
| 460 | Cha-F-100064 | II | 0.64 |
| 598 | VGlut-F-100285 | V | 0.58 |
| 935 | VGlut-F-100111 | V | 0.54 |
| 776 | VGlut-F-500423 | IV | 0.53 |
Cluster composition
This cluster has 24 neurons. The exemplar of this cluster (VGlut-F-000074) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| VGlut-F-000074 | 16005 | DvGlutMARCM-F394_seg1 | VGlut-Gal4 | F | 12573.71 | 1.00 |
| Cha-F-700190 | 3879 | ChaMARCM-F001545_seg001 | Cha-Gal4 | F | 8068.30 | 0.53 |
| Gad1-F-900062 | 4044 | GadMARCM-F000645_seg001 | Gad1-Gal4 | F | 6050.08 | 0.37 |
| Cha-F-100262 | 4182 | ChaMARCM-F001287_seg002 | Cha-Gal4 | F | 3024.00 | 0.26 |
| Cha-F-000371 | 4459 | ChaMARCM-F001501_seg001 | Cha-Gal4 | F | 8677.90 | 0.47 |
| VGlut-F-600391 | 6038 | DvGlutMARCM-F002707_seg002 | VGlut-Gal4 | F | 7422.87 | 0.62 |
| Cha-F-400110 | 7659 | ChaMARCM-F000489_seg001 | Cha-Gal4 | F | 8401.49 | 0.42 |
| Cha-F-200098 | 7665 | ChaMARCM-F000492_seg001 | Cha-Gal4 | F | 6284.55 | 0.40 |
| Cha-F-500052 | 8144 | ChaMARCM-F000179_seg002 | Cha-Gal4 | F | 7804.81 | 0.59 |
| VGlut-F-400727 | 8721 | DvGlutMARCM-F003553_seg001 | VGlut-Gal4 | F | 7424.23 | 0.50 |
| VGlut-F-600651 | 8963 | DvGlutMARCM-F003521_seg003 | VGlut-Gal4 | F | 9326.72 | 0.76 |
| VGlut-F-900079 | 9134 | DvGlutMARCM-F003427_seg002 | VGlut-Gal4 | F | 8867.54 | 0.72 |
| Gad1-F-800008 | 10606 | GadMARCM-F000115_seg002 | Gad1-Gal4 | F | 3875.61 | 0.45 |
| VGlut-F-700048 | 10820 | DvGlutMARCM-F2117_seg1 | VGlut-Gal4 | F | 7855.72 | 0.53 |
| VGlut-F-400562 | 12018 | DvGlutMARCM-F002538_seg002 | VGlut-Gal4 | F | 6709.09 | 0.44 |
| VGlut-F-400431 | 12769 | DvGlutMARCM-F1823_seg1 | VGlut-Gal4 | F | 7686.98 | 0.56 |
| VGlut-F-000444 | 13080 | DvGlutMARCM-F2102_seg2 | VGlut-Gal4 | F | 8088.79 | 0.53 |
| VGlut-F-200152 | 14506 | DvGlutMARCM-F779_seg1 | VGlut-Gal4 | F | 6405.09 | 0.44 |
| VGlut-F-400403 | 15319 | DvGlutMARCM-F1659_seg1 | VGlut-Gal4 | F | 6663.88 | 0.52 |
| VGlut-F-500001 | 15595 | DvGlutMARCM-F011_seg2 | VGlut-Gal4 | F | 7867.91 | 0.52 |
| VGlut-F-800003 | 15703 | DvGlutMARCM-F119-X2_seg2 | VGlut-Gal4 | F | 8377.23 | 0.63 |
| VGlut-F-000086 | 16036 | DvGlutMARCM-F427_seg1 | VGlut-Gal4 | F | 7306.61 | 0.51 |
| VGlut-F-000094 | 16076 | DvGlutMARCM-F461_seg1 | VGlut-Gal4 | F | 7149.32 | 0.53 |
| VGlut-F-300096 | 16226 | DvGlutMARCM-F600-X2_seg2 | VGlut-Gal4 | F | 5552.97 | 0.42 |