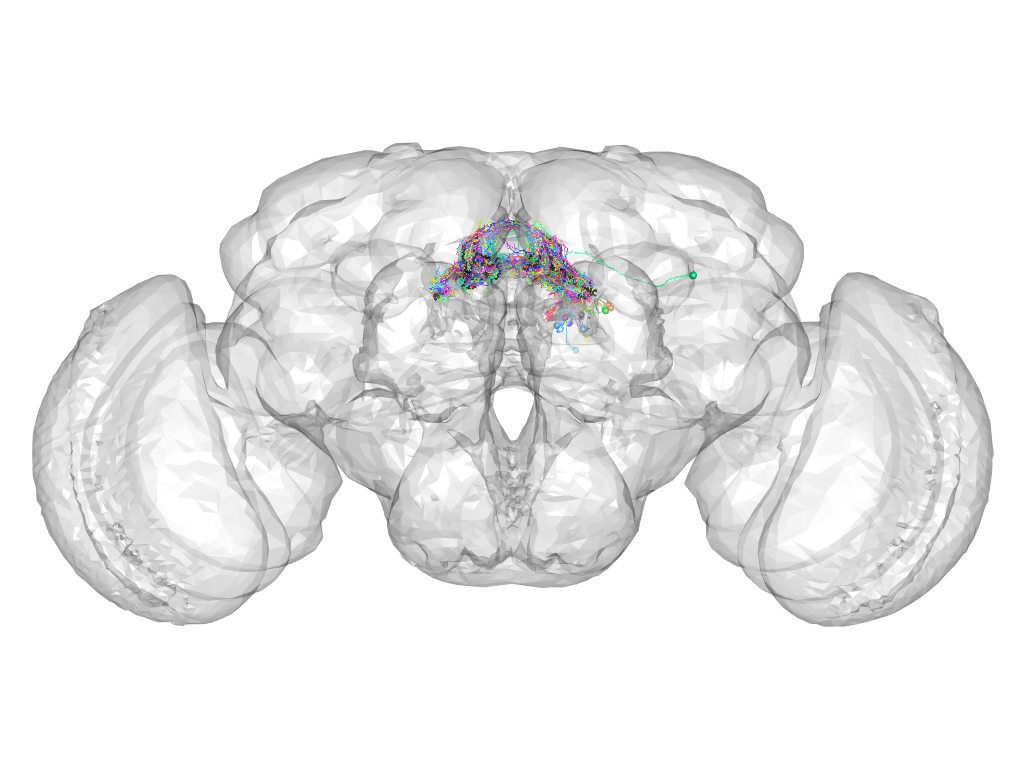
Cluster 1018
Part of supercluster XIV
About
Scores are NBLAST (version 2) similarity scores between each neuron and the exemplar (VGlut-F-300036), normalised to the self-score of the exemplar. The higher the better! Scores > 0 indicate a match of some sort. Excellent matches will have scores > 0.5.
Do you recognise any of the neurons in this cluster? Email us!
Credits
Raw data provided by flycircuit.tw as described in Three-Dimensional Reconstruction of Brain-wide Wiring Networks in Drosophila at Single-Cell Resolution by Ann-Shyn Chiang et al.
Image processing carried out with unu, CMTK, and Fiji.
Further processing carried out in R with nat. Clustering used package apcluster. 3D visualisation written from rgl by writeWebGL.
Similar clusters
The most similar clusters to this one are:
| Cluster | Exemplar | Supercluster | Exemplar normalised score |
|---|---|---|---|
| 1028 | VGlut-F-000067 | XIV | 0.70 |
| 964 | VGlut-F-200325 | XIV | 0.67 |
| 192 | TH-M-300027 | XIV | 0.55 |
| 142 | Trh-M-400110 | XIV | 0.54 |
| 204 | Trh-M-100025 | XIV | 0.50 |
Cluster composition
This cluster has 57 neurons. The exemplar of this cluster (VGlut-F-300036) is drawn in black.
| Neuron | idid | Gene name | Driver | Sex | Raw score | Normalised score |
|---|---|---|---|---|---|---|
| VGlut-F-300036 | 15912 | DvGlutMARCM-F310-X2_seg1 | VGlut-Gal4 | F | 35397.73 | 1.00 |
| VGlut-F-500843 | 5741 | DvGlutMARCM-F004448_seg001 | VGlut-Gal4 | F | 24391.89 | 0.75 |
| VGlut-F-500845 | 5746 | DvGlutMARCM-F004451_seg002 | VGlut-Gal4 | F | 23701.35 | 0.72 |
| VGlut-F-700574 | 5773 | DvGlutMARCM-F004477_seg001 | VGlut-Gal4 | F | 24481.59 | 0.73 |
| VGlut-F-700603 | 5942 | DvGlutMARCM-F004598_seg003 | VGlut-Gal4 | F | 25024.00 | 0.74 |
| VGlut-F-800328 | 5956 | DvGlutMARCM-F004608_seg001 | VGlut-Gal4 | F | 23790.09 | 0.71 |
| VGlut-F-600134 | 6097 | DvGlutMARCM-F2153_seg1 | VGlut-Gal4 | F | 24798.55 | 0.75 |
| VGlut-F-400883 | 6456 | DvGlutMARCM-F004241_seg001 | VGlut-Gal4 | F | 23647.75 | 0.72 |
| VGlut-F-400886 | 6460 | DvGlutMARCM-F004245_seg001 | VGlut-Gal4 | F | 26087.81 | 0.77 |
| VGlut-F-500795 | 6489 | DvGlutMARCM-F004267_seg001 | VGlut-Gal4 | F | 23456.52 | 0.71 |
| VGlut-F-800272 | 6518 | DvGlutMARCM-F004291_seg001 | VGlut-Gal4 | F | 24312.69 | 0.73 |
| VGlut-F-700537 | 6570 | DvGlutMARCM-F004332_seg001 | VGlut-Gal4 | F | 24040.22 | 0.73 |
| VGlut-F-700442 | 6704 | DvGlutMARCM-F003918_seg001 | VGlut-Gal4 | F | 22457.52 | 0.71 |
| VGlut-F-400810 | 6730 | DvGlutMARCM-F003938_seg002 | VGlut-Gal4 | F | 24102.89 | 0.70 |
| VGlut-F-400811 | 6731 | DvGlutMARCM-F003939_seg001 | VGlut-Gal4 | F | 24490.72 | 0.71 |
| VGlut-F-400817 | 6738 | DvGlutMARCM-F003943_seg001 | VGlut-Gal4 | F | 24390.44 | 0.74 |
| VGlut-F-500741 | 6739 | DvGlutMARCM-F003944_seg001 | VGlut-Gal4 | F | 24530.86 | 0.74 |
| VGlut-F-700459 | 6762 | DvGlutMARCM-F003965_seg001 | VGlut-Gal4 | F | 24386.55 | 0.73 |
| VGlut-F-700470 | 6803 | DvGlutMARCM-F003993_seg001 | VGlut-Gal4 | F | 24505.31 | 0.74 |
| VGlut-F-500753 | 6808 | DvGlutMARCM-F003998_seg002 | VGlut-Gal4 | F | 24829.55 | 0.73 |
| VGlut-F-400828 | 6849 | DvGlutMARCM-F004026_seg001 | VGlut-Gal4 | F | 24039.89 | 0.72 |
| VGlut-F-700477 | 6932 | DvGlutMARCM-F004087_seg001 | VGlut-Gal4 | F | 24793.12 | 0.72 |
| VGlut-F-700493 | 7026 | DvGlutMARCM-F004163_seg001 | VGlut-Gal4 | F | 23171.03 | 0.70 |
| VGlut-F-700508 | 7050 | DvGlutMARCM-F004180_seg001 | VGlut-Gal4 | F | 23337.05 | 0.71 |
| VGlut-F-500777 | 7069 | DvGlutMARCM-F004199_seg001 | VGlut-Gal4 | F | 24597.62 | 0.74 |
| fru-F-400126 | 7365 | FruMARCM-F000993_seg001 | fru-Gal4 | F | 18983.79 | 0.54 |
| VGlut-F-400788 | 7402 | DvGlutMARCM-F003860_seg001 | VGlut-Gal4 | F | 24830.30 | 0.74 |
| VGlut-F-600692 | 7405 | DvGlutMARCM-F003862_seg002 | VGlut-Gal4 | F | 24727.87 | 0.74 |
| Gad1-F-700005 | 8401 | GadMARCM-F000130_seg001 | Gad1-Gal4 | F | 23623.56 | 0.71 |
| VGlut-F-700277 | 8514 | DvGlutMARCM-F003096_seg003 | VGlut-Gal4 | F | 25260.51 | 0.74 |
| VGlut-F-700295 | 8538 | DvGlutMARCM-F003114_seg001 | VGlut-Gal4 | F | 24928.92 | 0.74 |
| VGlut-F-700404 | 8618 | DvGlutMARCM-F003672_seg003 | VGlut-Gal4 | F | 24154.27 | 0.72 |
| VGlut-F-700356 | 8895 | DvGlutMARCM-F003473_seg002 | VGlut-Gal4 | F | 25604.48 | 0.74 |
| VGlut-F-700320 | 9254 | DvGlutMARCM-F003315_seg001 | VGlut-Gal4 | F | 23212.90 | 0.70 |
| VGlut-F-300481 | 9364 | DvGlutMARCM-F003197_seg001 | VGlut-Gal4 | F | 25243.82 | 0.75 |
| Gad1-F-100041 | 9588 | GadMARCM-F000457_seg001 | Gad1-Gal4 | F | 24667.05 | 0.73 |
| Gad1-F-600011 | 10585 | GadMARCM-F000103_seg001 | Gad1-Gal4 | F | 23630.19 | 0.71 |
| VGlut-F-600403 | 10780 | DvGlutMARCM-F002730_seg001 | VGlut-Gal4 | F | 24238.13 | 0.73 |
| VGlut-F-700145 | 10870 | DvGlutMARCM-F002782_seg003 | VGlut-Gal4 | F | 26430.23 | 0.76 |
| VGlut-F-600464 | 11019 | DvGlutMARCM-F002902_seg001 | VGlut-Gal4 | F | 23956.31 | 0.73 |
| VGlut-F-600505 | 11660 | DvGlutMARCM-F002234_seg001 | VGlut-Gal4 | F | 25591.79 | 0.75 |
| VGlut-F-600169 | 11730 | DvGlutMARCM-F002303_seg001 | VGlut-Gal4 | F | 24823.47 | 0.75 |
| VGlut-F-600187 | 11749 | DvGlutMARCM-F002321_seg001 | VGlut-Gal4 | F | 23635.46 | 0.72 |
| VGlut-F-600192 | 11754 | DvGlutMARCM-F002325_seg001 | VGlut-Gal4 | F | 24993.51 | 0.74 |
| VGlut-F-500397 | 11964 | DvGlutMARCM-F002498_seg001 | VGlut-Gal4 | F | 24558.50 | 0.74 |
| VGlut-F-600388 | 12243 | DvGlutMARCM-F002705_seg001 | VGlut-Gal4 | F | 25744.03 | 0.74 |
| VGlut-F-400579 | 12285 | DvGlutMARCM-F002741_seg001 | VGlut-Gal4 | F | 22401.84 | 0.68 |
| VGlut-F-400446 | 12830 | DvGlutMARCM-F1877_seg2 | VGlut-Gal4 | F | 25135.40 | 0.75 |
| VGlut-F-500330 | 13036 | DvGlutMARCM-F2060_seg2 | VGlut-Gal4 | F | 25830.64 | 0.75 |
| VGlut-F-500364 | 13129 | DvGlutMARCM-F2156_seg1 | VGlut-Gal4 | F | 24769.54 | 0.74 |
| VGlut-F-400302 | 14373 | DvGlutMARCM-F1053_seg1 | VGlut-Gal4 | F | 24408.08 | 0.74 |
| VGlut-F-400192 | 14459 | DvGlutMARCM-F745-X2_seg1 | VGlut-Gal4 | F | 24951.71 | 0.75 |
| VGlut-F-400198 | 14469 | DvGlutMARCM-F749-x2_seg2 | VGlut-Gal4 | F | 25278.00 | 0.75 |
| VGlut-F-300169 | 14540 | DvGlutMARCM-F809_seg1 | VGlut-Gal4 | F | 24111.51 | 0.71 |
| VGlut-F-400310 | 14748 | DvGlutMARCM-F1100_seg1 | VGlut-Gal4 | F | 24748.21 | 0.75 |
| VGlut-F-500177 | 15174 | DvGlutMARCM-F1505_seg1 | VGlut-Gal4 | F | 25170.19 | 0.74 |
| VGlut-F-400063 | 15951 | DvGlutMARCM-F346_seg1 | VGlut-Gal4 | F | 24843.52 | 0.68 |